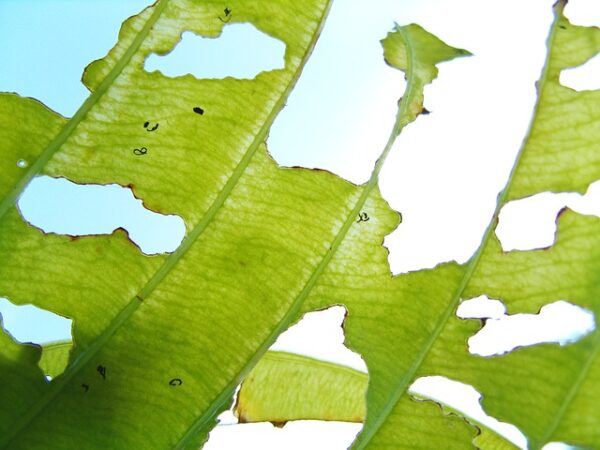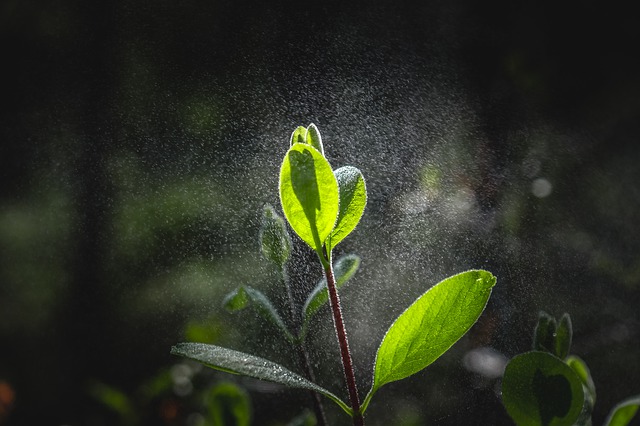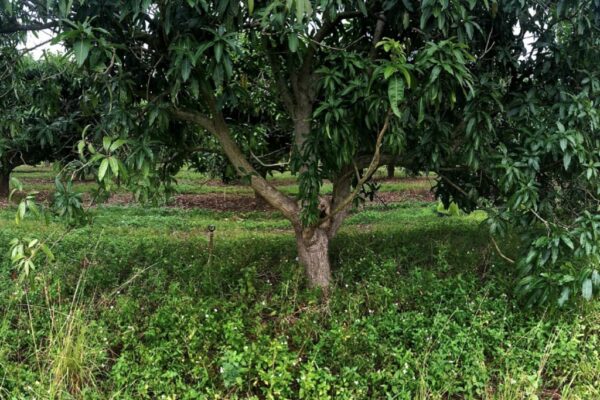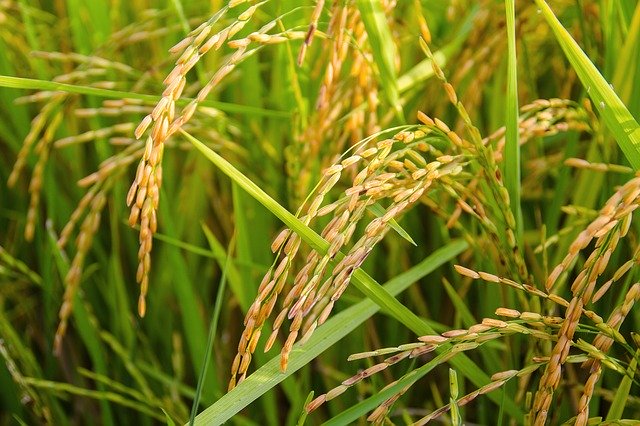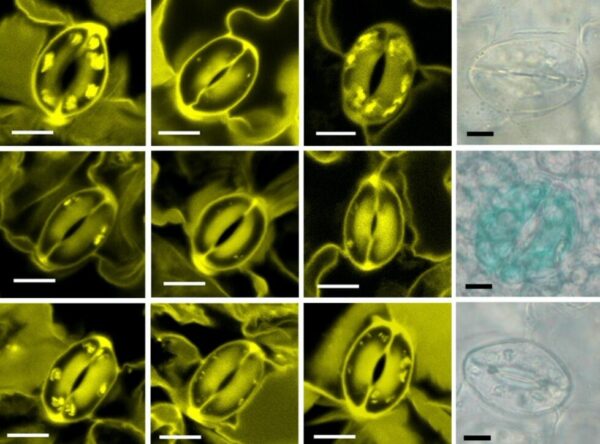
Wildfires are devastating, but they can also bring new life by clearing existing vegetation and allowing new plants to spring up. Many plants in fire-prone areas actually require exposure to fire for seeds to germinate. In the past decade, scientists…
Read More