This week we spoke to Professor Jonathan Lynch, Penn State University, whose research on root traits has deepened our understanding of how plants adapt to drought and low soil fertility.
This week we spoke to Professor Jonathan Lynch, Penn State University, whose research on root traits has deepened our understanding of how plants adapt to drought and low soil fertility.
Could you begin by giving us a brief introduction to your research?
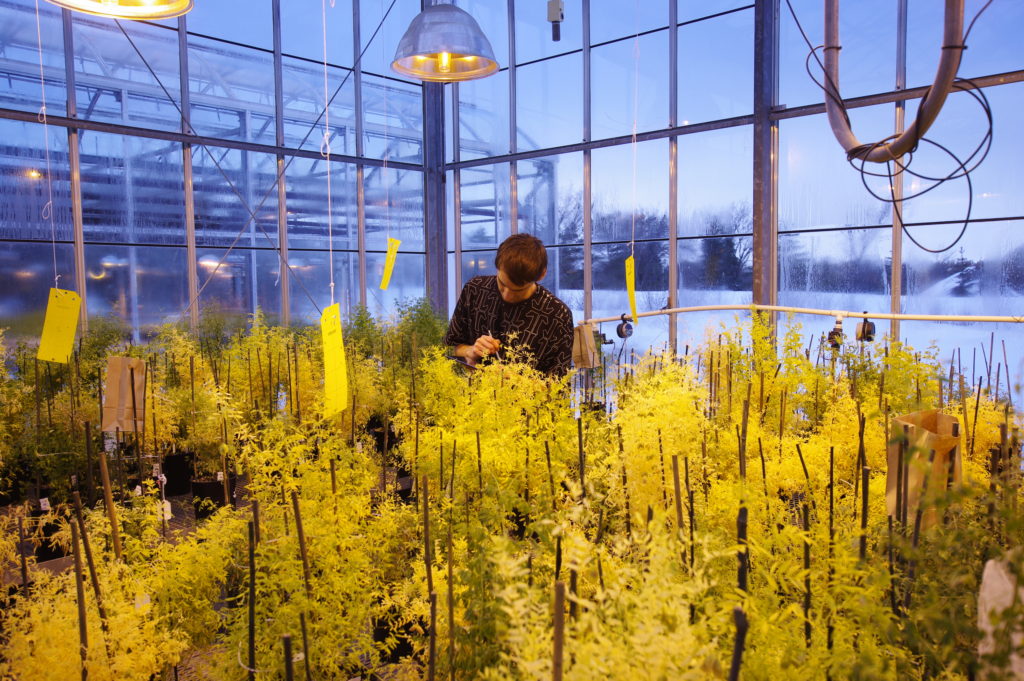
We are trying to understand how plants adapt to drought and low soil fertility. This is important because all plants in terrestrial ecosystems experience suboptimal water and nutrient availability, so in rich nations we maintain crop yields with irrigation and fertilizer, which is not sustainable in the long term. Furthermore, climate change is further degrading soil fertility and increasing plant stress. This topic is therefore both a central question in plant evolution and a key challenge for our civilization. We need to develop better ways to sustain so many people on this planet, and a big part of that will be developing more resilient, efficient crop plants.
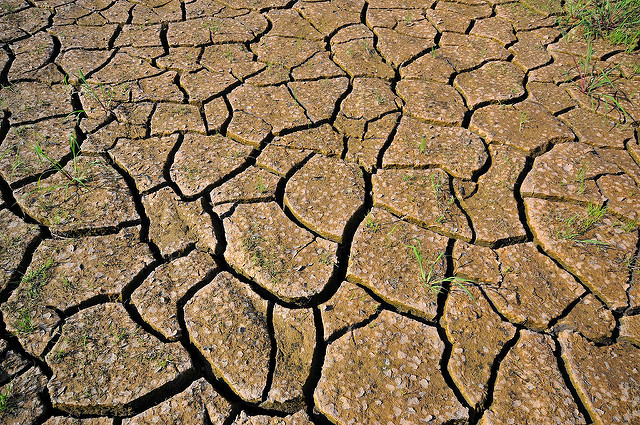

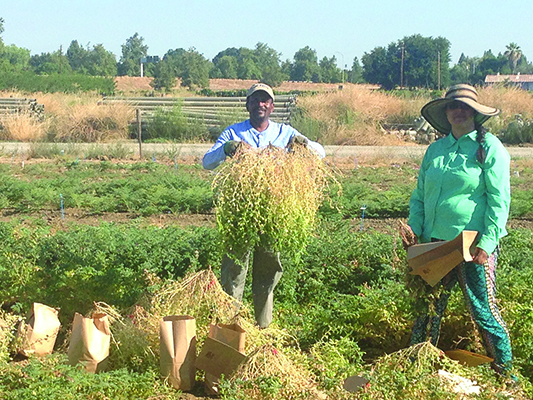
Drought and low soil fertility are devastating for crops. Image credit: CIAT. Used under license: CC BY-SA 2.0.
What got you interested in this field, and how has your career developed over time?
When I was 9 years old I became aware of a famine in Africa related to crop failure and resolved to do something about it. I studied soils and plant nutrition as an undergraduate, and in graduate school worked on plant adaptation to low phosphorus and salinity stress, moving to a research position at the CIAT headquarters in Colombia. Later I moved to Penn State, where I have maintained this focus, working to understand the stress tolerance of staple crops, and collaborating with crop breeders in the USA, Europe, Africa, Asia, and Latin America.
Your recent publications feature a variety of different crop plants. Could you talk about how you select a species to study?
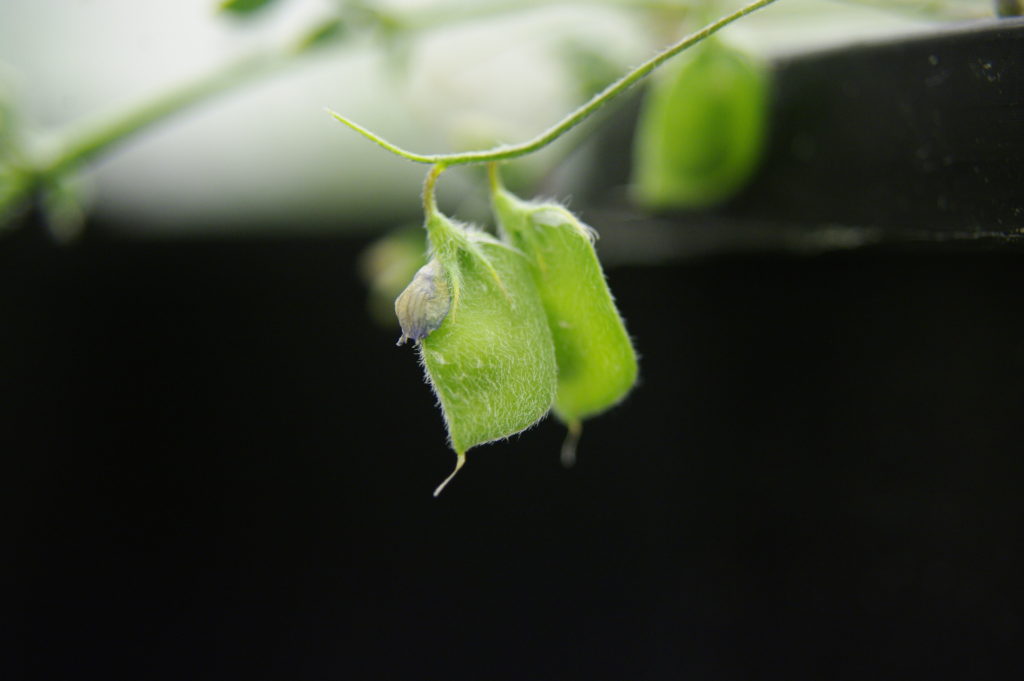
We work with species that are important for food security, that grow in our field environments, and that I think are cool. We have devoted most of our efforts to the common bean – globally the most important food legume – and maize, which is the most important global crop. These species are often grown together in Africa and Latin America, and part of our work has been geared to understanding how maize/bean and maize/bean/squash polycultures perform under stress. These are fascinating, beautiful plants with huge cultural importance in human history. They are also supported by talented, cooperative research communities. One nice feature of working with food security crops is that their research communities share common goals of achieving impact to improve human welfare.
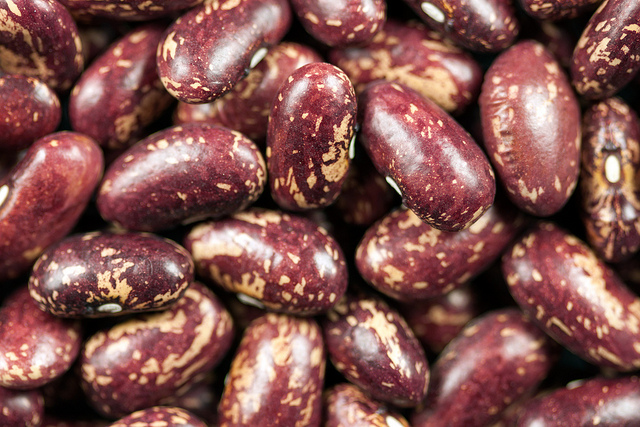

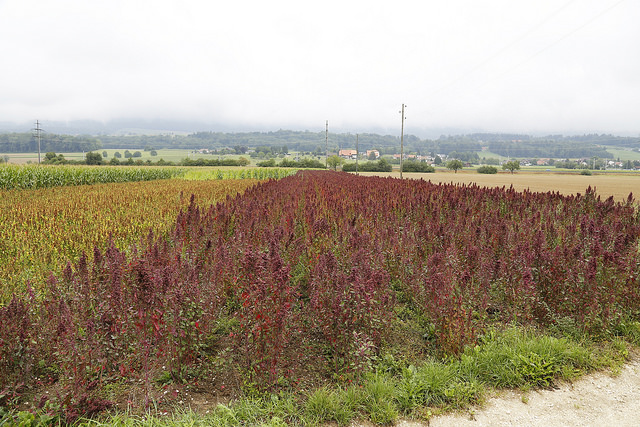
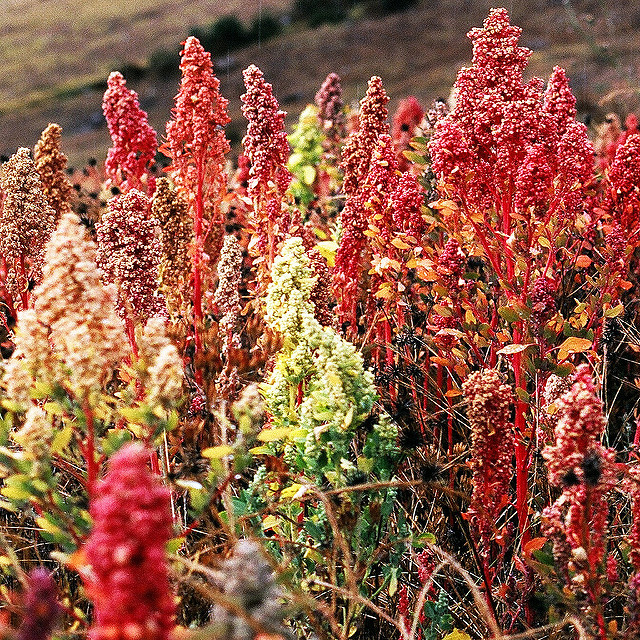
The common bean (Phaseolus vulgaris) is an important staple in many parts of the world. Image credit: Ervins Strauhmanis. Used under license: CC BY 2.0.
Many researchers use Arabidopsis thaliana for plant research, but are crops better suited for root research than the delicate roots of Arabidopsis? Are crop plants more or less difficult to work with in your research than Arabidopsis?
The best research system is entirely a function of your goals and questions. We have worked with Arabidopsis for some questions. Since we work with processes at multiple scales, including crop stands, whole organisms, organs, tissues, and cells, it has been useful to work with large plants such as maize, which are large enough to easily measure and to work with in the field. The most interesting stress adaptations for crop breeding are those that differ among genotypes of the same species, and at that level of organization there is a lot of biology that is specific to that species, that cannot readily be generalized from model organisms with very different life strategies. There has been considerable attention to model genomes and much less attention to model phenomes.
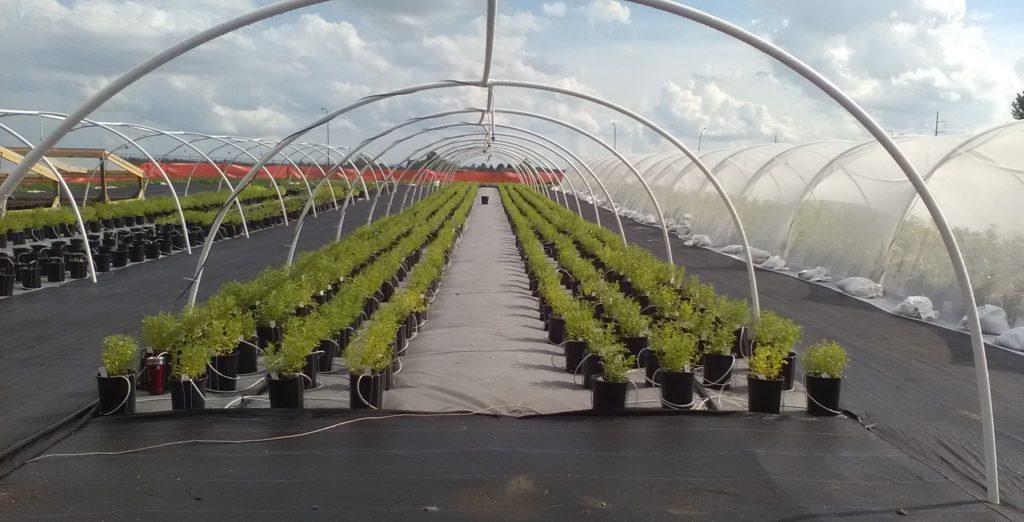
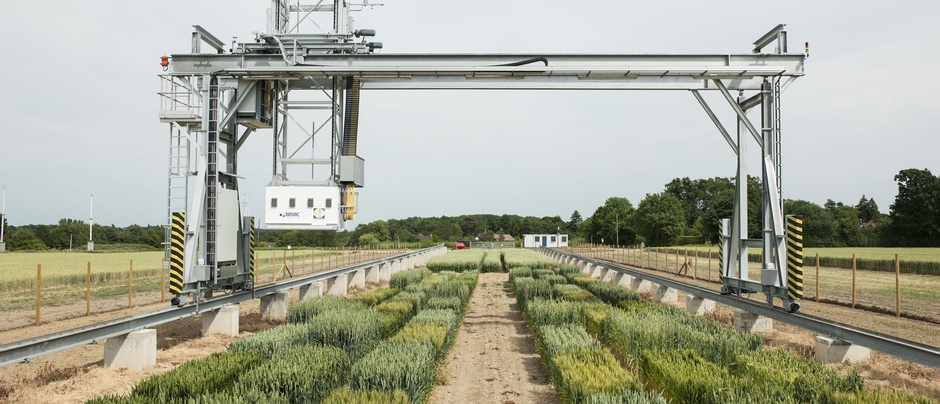
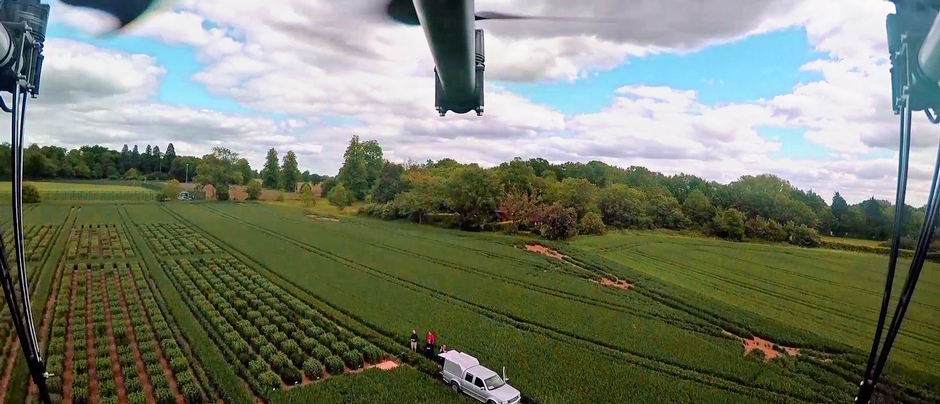
You have developed methodologies for the high-throughput phenotyping of crop plants. What does this technique involve and what challenges did you have to overcome to succeed?
We have developed multiple phenotyping approaches – too many to summarize readily here. Our overall approach is simply to develop a tool that helps us achieve our goals. For example, we have developed tools to quantify the root architecture of thousands of plants in the field, to measure anatomical phenotypes of thousands of samples from field-grown roots, to help us determine which root phenotypes might affect soil resource capture, etc. Working with geneticists and breeders, we are constantly asked to measure something meaningful on thousands of plants in a field, in many fields, every season. ARPA-E (the US Advanced Research Projects Agency for Energy) has recently funded us to develop phenotyping tools for root depth in the field, but this is the first time we have been funded to develop phenotyping tools – generally we just come up with things to help us do our work, which fortunately have been useful for other researchers as well.
Could you talk about some of the computational models you have developed for investigating plant growth and development?
The biological interactions between plants and their environment are so complex, we need computational (in silico) tools to help us evaluate them. Increasingly, in silico tools can integrate information across multiple scales, from gene expression to crop stands. These tools also allow us to evaluate things that are difficult to measure, such as phenotypes that do not yet exist, or future climates. In silico biology will be an essential tool in 21st Century biology, which will have access to huge amounts of data at multiple scales that can be used to try to understand incredibly complex systems, such as the human brain or roots interacting with living soil. Our main in silico tool is SimRoot, developed over the past 25 years to understand how root phenotypes affect soil resource capture.
Check out a SimRoot model below:
You have been working on breeding plants that have improved yield in soils with low fertility. What have you achieved in this work?
In collaboration with crop breeders and colleagues in various nations we have developed improved common bean lines with better yield under drought and low soil fertility that are being deployed in Africa and Latin America, improved soybean lines with better yield in soils with low phosphorus being deployed in Africa and Asia, and are now working with maize breeders in Africa to develop lines with better yield under drought and low nitrogen stress. Many crop breeders are using our methods for root phenotyping to target root phenotypes in their selection regimes in multiple crops.
What piece of advice do you have for early career researchers?
You are at the forefront of an unprecedented challenge we face as a species – how to sustain 10 billion people in a degrading environment. Plant biologists are an essential part of the effort to reshape how we live on this planet. Do not doubt the importance of your efforts. Do not lose sight of the very real human impact of your scientific choices. Do not be deterred by the gamesmanship and ‘primate politics’ of science. You can make a difference. We need you.