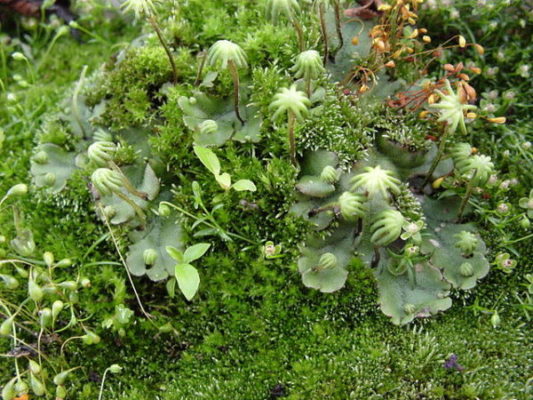
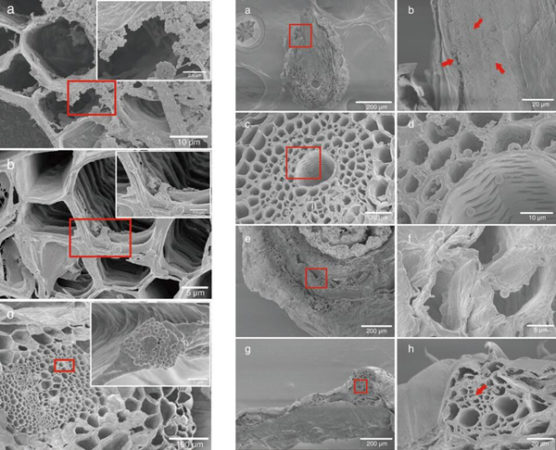
An international study has discovered a stem-cell promoting hormone in the liverwort Marchantia polymopha. Marchantia, a common liverwort, is a representative of an ancient lineage of plants. Their evolutionary history presents researchers with an excellent opportunity to explore the fundamental…
Read More