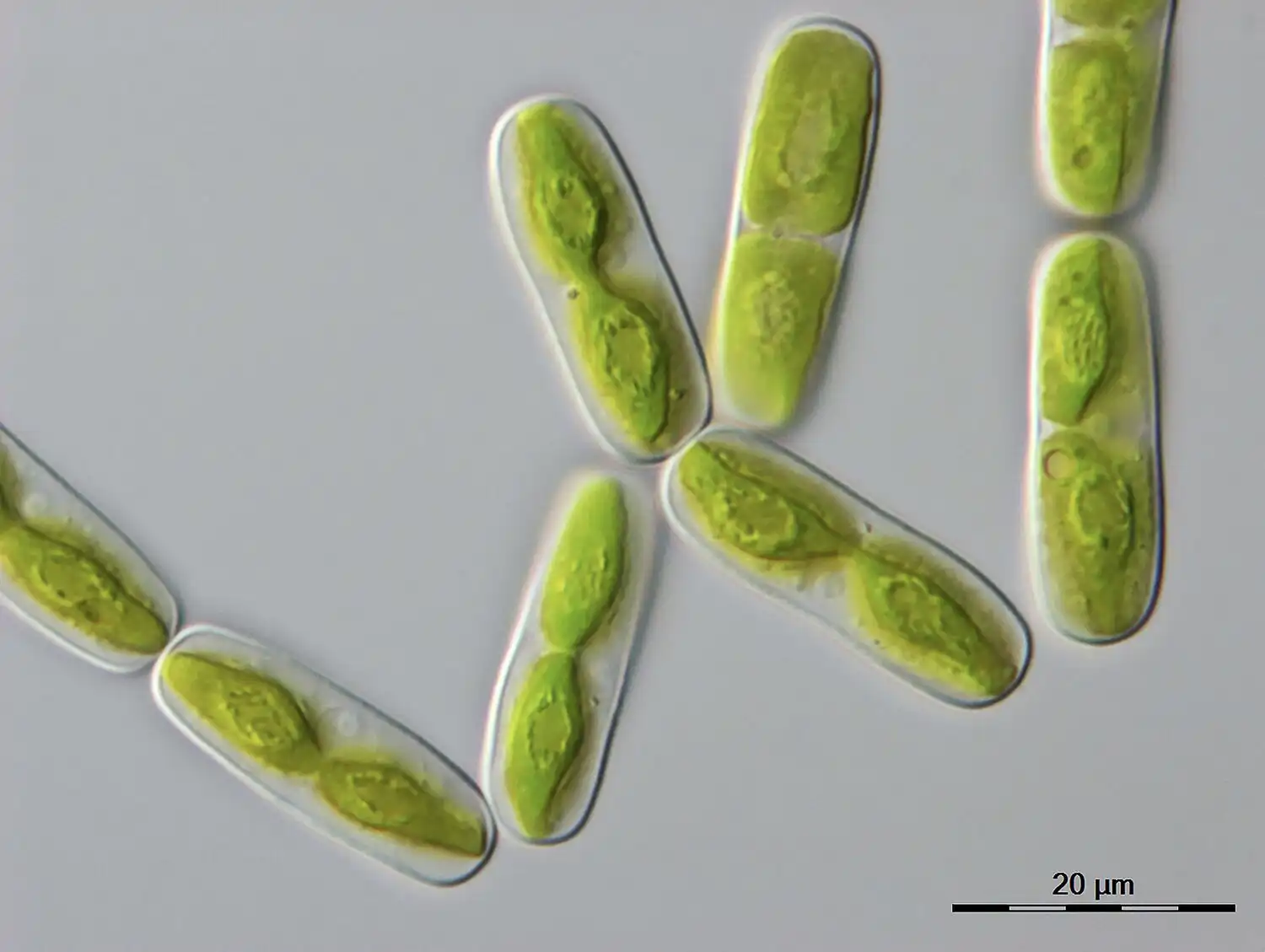
A long-lived monocarpic species of bamboo, Phyllostachys nigra var. henonis, only flowers once every 120 years before it dies. The upcoming flowering event for this species does not bode well for its continued long-term survival, as most flowers are not producing…
Read More