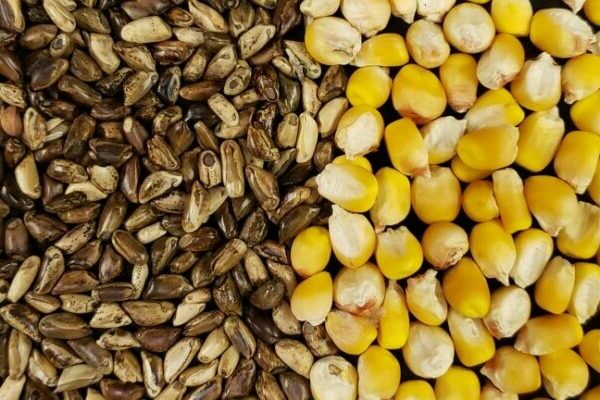
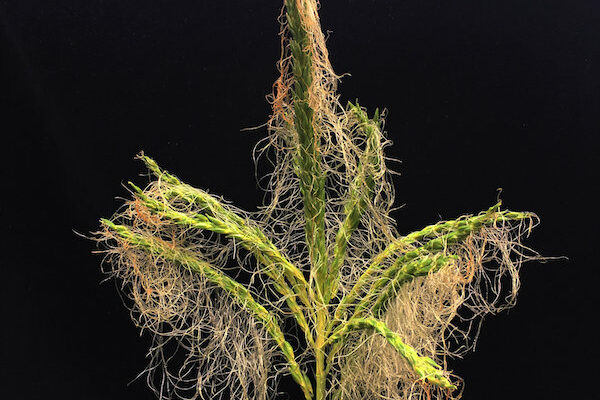
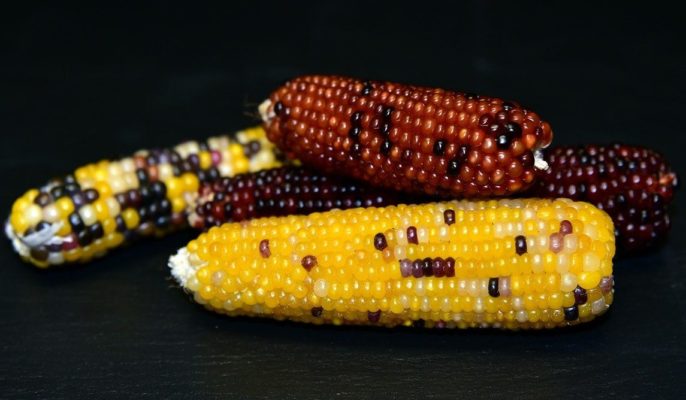
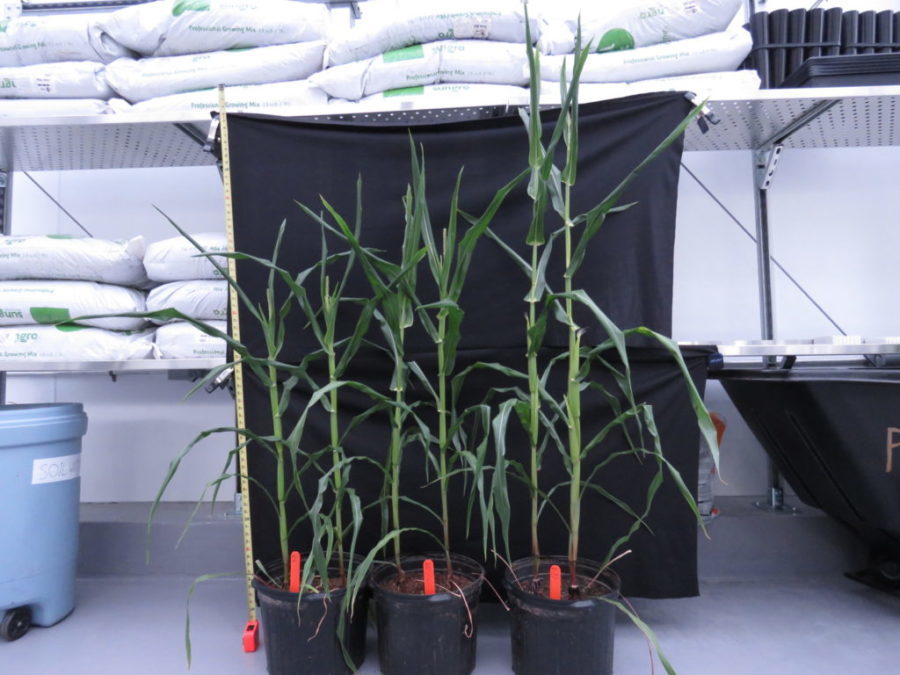
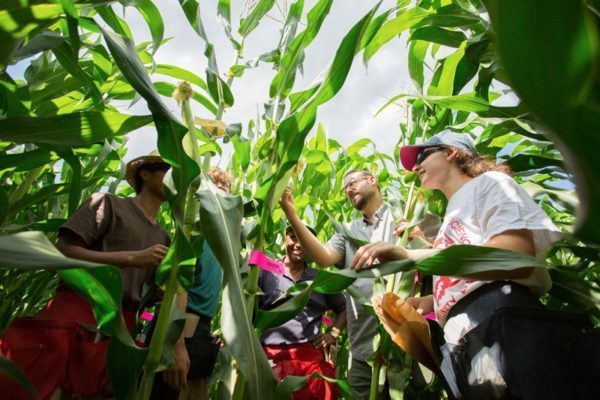









Maize is a staple food all over the world. In the United States, where it’s better known as corn, nearly 90 million acres were planted in 2018, earning $47.2 billion in crop cash receipts.
But, under the effects of climate change, this signature crop may not fare so well. As the world tries to feed a population skyrocketing to nine billion by 2050, that has major implications. So, what can we do about it? The answer might be exotic.
A multi-institutional team led by University of Delaware plant geneticist Randy Wisser decoded the genetic map for how maize from tropical environments can be adapted to the temperate U.S. summer growing season. Wisser sees these exotic varieties, which are rarely used in breeding, as key to creating next-era varieties of corn.
The research team included scientists from UD, North Carolina State University, University of Wisconsin, University of Missouri, Iowa State University, Texas A&M University and the U.S. Department of Agriculture-Agricultural Research Service. The resulting study, highlighted by the editorial board of Genetics, provides a new lens into the future viability of one of the world’s most important grains.
“If we can expand the genetic base by using exotic varieties, perhaps we can counter stresses such as emerging diseases and drought associated with growing corn in a changing climate,” said Wisser, associate professor in UD’s Department of Plant and Soil Sciences. “That is critical to ensuring ample production for the billions of people who depend on it for food and other products.”
Modern maize strains were created from only a small fraction of the global maize population. This limited infusion of diversity raises concerns about the vulnerability of American corn in a shifting climate. The U.S. Department of Agriculture (USDA) seed bank includes tens of thousands of varieties, but many are just not being used.
“We know that the tropical maize varieties represent our greatest reservoir of genetic diversity,” said study co-author Jim Holland, a plant geneticist with the USDA Agricultural Research Service at North Carolina State. “This study improved our understanding of those genetics, so we can use this information to guide future breeding efforts to safeguard the corn crop.”
Certain exotic strains of maize better handle drought or waterlogging or low-nitrogen soil, for example. But because these strains have evolved outside the U.S., they are not immediately suited to states like Delaware. So, exotics first need to be pre-adapted.
In prior work, Wisser and his colleagues showed how 10 years of repeated genetic selection was required to adapt a tropical strain of maize to the temperate U.S. Co-author Arnel Hallauer spent a decade adapting the population through selective breeding, so it could flourish in an environment like Delaware.
“What’s so cool now is that we could go back to the original generations from Dr. Hallauer and grow them side by side in the same field,” Wisser said of the first-of-its-kind experimental design. “This allows us to rule out the influence of the environment on each trait, directly exposing the genetic component of evolution. This has opened a ‘back to the future’ channel where we can redesign our approach to developing modern varieties.”
While extremely impressive, a decade to adapt exotic maize to new environments is a lot of time when the climate change clock is ticking.
“Unfortunately, this process takes 10 years, which is not counting ongoing evaluations and integrating the exotic variations into more commonly used types of maize,” Wisser said. “With the climate threats we face, that’s a long time. So, gaining insights into this evolutionary process will help us devise ways to shorten the time span.”
Accelerating adaptation
Wisser isn’t wasting any time as he explores ways to bolster corn’s ability to survive and thrive. He and Holland are working on a new project to cut that time span in half.
In cutting-edge research funded by the U.S. Department of Agriculture’s National Institute of Food and Agriculture, the team is analyzing how corn genomes behave in a target environment as they aim to formulate a predictive model for fitness.
“What we’re doing is sequencing the genomes and measuring traits like flowering time or disease for individuals in one generation. From this, we can generate a lookup table that allows us to foresee which individuals in the next generation have the best traits based on their genetic profiles alone,” Wisser said. “And our lookup table can be tailored to predict how the individuals will behave in a particular environment or location like Delaware.”
That means plant breeders could grow a second generation of corn anywhere outside of Delaware, but still predict which individuals would be the most fit for Delaware’s environment.
“For instance, even if the plants are grown at a location where a disease is not present, our prediction model can still select the resistant plants and cross them to enrich the genes that underlie resistance,” Wisser said.
With this approach, researchers don’t have to wait out a Delaware winter, so they can continue to pre-adapt the population for at least one extra generation per year. That’s how 10 years of selective breeding for pre-adaptation could become five, providing a quicker route to access exotic genes.
This new effort connects to the Genomes To Fields (G2F) Initiative, developed in 2013 for understanding and capitalizing on the link between genomes and crop performance for the benefit of growers, consumers and society.
If Wisser and Holland can develop a method to rapidly pre-adapt exotics, this opens a lane for G2F to test the impact of these unique genomes on crop performance.
“Our goal is to advance the science so breeders can draw on a wider array of the diversity that has accumulated across thousands of years of evolution,” explained Wisser, who has been involved in the public-private initiative since its beginning. “In turn, they can produce improved varieties for producers and consumers facing the challenges of climate change.”
Read the paper: Genetics
Article source: University of Delaware
Author: Dante LaPenta
Image credit: Monica Moriak, Evan Krape, Teclemariam Weldekidan and Randy Wisser
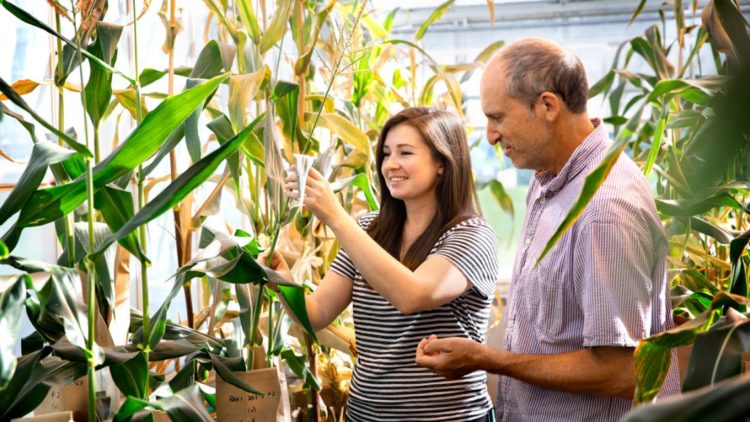

Nearly everyone on Earth is familiar with corn. Literally.
Around the world, each person eats an average of 70 pounds of the grain each year, with even more grown for animal feed and biofuel. And as the global population continues to boom, increasing the amount of food grown on the same amount of land becomes increasingly important.
One potential solution is to develop crops that perform better in cold temperatures. Many people aren’t aware that corn is a tropical plant, which makes it extremely sensitive to cold weather. This trait is problematic in temperate climates where the growing season averages only 4 or 5 months – and where more than 60% of its 1.6 trillion pound annual production occurs.
A chilling-tolerant strain could broaden the latitudes in which the crop could be grown, as well as enable current farmers to increase productivity.
A group of researchers led by David Stern, president of the Boyce Thompson Institute, have taken a step closer to this goal by developing a new type of corn that recovers much more quickly after a cold snap. Stern is also an adjunct professor of plant biology in Cornell University’s College of Agriculture and Life Sciences.
The research is described in a paper published online in Plant Biotechnology Journal.
“In the field, chilling stress happens most often in the spring when cold temperatures combine with strong sunlight, causing plants to bleach,” Stern said. “So a more chilling-tolerant corn could help farmers plant earlier in the year with confidence that their crop would survive a cold spell and bounce back quickly once the weather warmed up again.”
This work built on research published in 2018, which showed that increasing levels of an enzyme called Rubisco led to bigger and faster-maturing plants. Rubisco is essential for plants to turn atmospheric carbon dioxide into sugar, and its levels in corn leaves decrease dramatically in cold weather.
In the latest study, Stern and colleagues grew corn plants for three weeks at 25°C (77°F), lowered the temperature to 14°C (57°F) for two weeks, and then increased it back up to 25°C.
“The corn with more Rubisco performed better than regular corn before, during and after chilling,” said Coralie Salesse-Smith, the paper’s first author. “In essence, we were able to reduce the severity of chilling stress and allow for a more rapid recovery.” Salesse-Smith was a Cornell PhD candidate in Stern’s lab during the study, and she is now a postdoctoral researcher at the University of Illinois.
Indeed, compared to regular corn, the engineered corn had higher photosynthesis rates throughout the experiment, and recovered more quickly from the chilling stress with less damage to the molecules that perform the light-dependent reactions of photosynthesis.
The end result was a plant that grew taller and developed mature ears of corn more quickly following a cold spell.
Steve Reiners, a co-team leader for Cornell Cooperative Extension’s vegetable program, says that sweet corn is a major vegetable crop in New York, worth about $40-$60 million annually. He notes that many New York corn growers plant as soon as they can because an early crop commands the highest prices of the season.
“Many corn growers in New York plant early under protective plastic sheets to increase soil temperatures, which is expensive. Chilling-tolerant corn could allow farmers to remove that plastic sooner,” Reiners said. “This would expose the plants to additional sunlight, potentially enabling them to mature earlier in the season and get farmers those higher prices.”
Reiners, who was not involved in the study, is also a professor of horticulture at Cornell.
“The corn we developed isn’t yet completely optimized for chilling tolerance, so we are planning the next generation of modifications,” said Stern. “For example, it would be very interesting to add a chilling-tolerant version of a protein called PPDK into the corn and see if it performs even better.”
The researchers believe their approach could also be used in other crops that use the C4 photosynthetic pathway to fix carbon, such as sugar cane and sorghum.
Co-authors on the paper include researchers from The Australian National University in Canberra.
Read the paper: Plant Biotechnology Journal
Article source: Boyce Thompson Institute
Author: Aaron J. Bouchie
Image credit: Jason Koski/Brand Communications
 This week we spoke to Professor Jonathan Lynch, Penn State University, whose research on root traits has deepened our understanding of how plants adapt to drought and low soil fertility.
This week we spoke to Professor Jonathan Lynch, Penn State University, whose research on root traits has deepened our understanding of how plants adapt to drought and low soil fertility.
Could you begin by giving us a brief introduction to your research?
We are trying to understand how plants adapt to drought and low soil fertility. This is important because all plants in terrestrial ecosystems experience suboptimal water and nutrient availability, so in rich nations we maintain crop yields with irrigation and fertilizer, which is not sustainable in the long term. Furthermore, climate change is further degrading soil fertility and increasing plant stress. This topic is therefore both a central question in plant evolution and a key challenge for our civilization. We need to develop better ways to sustain so many people on this planet, and a big part of that will be developing more resilient, efficient crop plants.

Drought and low soil fertility are devastating for crops. Image credit: CIAT. Used under license: CC BY-SA 2.0.
What got you interested in this field, and how has your career developed over time?
When I was 9 years old I became aware of a famine in Africa related to crop failure and resolved to do something about it. I studied soils and plant nutrition as an undergraduate, and in graduate school worked on plant adaptation to low phosphorus and salinity stress, moving to a research position at the CIAT headquarters in Colombia. Later I moved to Penn State, where I have maintained this focus, working to understand the stress tolerance of staple crops, and collaborating with crop breeders in the USA, Europe, Africa, Asia, and Latin America.
Your recent publications feature a variety of different crop plants. Could you talk about how you select a species to study?
We work with species that are important for food security, that grow in our field environments, and that I think are cool. We have devoted most of our efforts to the common bean – globally the most important food legume – and maize, which is the most important global crop. These species are often grown together in Africa and Latin America, and part of our work has been geared to understanding how maize/bean and maize/bean/squash polycultures perform under stress. These are fascinating, beautiful plants with huge cultural importance in human history. They are also supported by talented, cooperative research communities. One nice feature of working with food security crops is that their research communities share common goals of achieving impact to improve human welfare.

The common bean (Phaseolus vulgaris) is an important staple in many parts of the world. Image credit: Ervins Strauhmanis. Used under license: CC BY 2.0.
Many researchers use Arabidopsis thaliana for plant research, but are crops better suited for root research than the delicate roots of Arabidopsis? Are crop plants more or less difficult to work with in your research than Arabidopsis?
The best research system is entirely a function of your goals and questions. We have worked with Arabidopsis for some questions. Since we work with processes at multiple scales, including crop stands, whole organisms, organs, tissues, and cells, it has been useful to work with large plants such as maize, which are large enough to easily measure and to work with in the field. The most interesting stress adaptations for crop breeding are those that differ among genotypes of the same species, and at that level of organization there is a lot of biology that is specific to that species, that cannot readily be generalized from model organisms with very different life strategies. There has been considerable attention to model genomes and much less attention to model phenomes.
You have developed methodologies for the high-throughput phenotyping of crop plants. What does this technique involve and what challenges did you have to overcome to succeed?
We have developed multiple phenotyping approaches – too many to summarize readily here. Our overall approach is simply to develop a tool that helps us achieve our goals. For example, we have developed tools to quantify the root architecture of thousands of plants in the field, to measure anatomical phenotypes of thousands of samples from field-grown roots, to help us determine which root phenotypes might affect soil resource capture, etc. Working with geneticists and breeders, we are constantly asked to measure something meaningful on thousands of plants in a field, in many fields, every season. ARPA-E (the US Advanced Research Projects Agency for Energy) has recently funded us to develop phenotyping tools for root depth in the field, but this is the first time we have been funded to develop phenotyping tools – generally we just come up with things to help us do our work, which fortunately have been useful for other researchers as well.
Could you talk about some of the computational models you have developed for investigating plant growth and development?
The biological interactions between plants and their environment are so complex, we need computational (in silico) tools to help us evaluate them. Increasingly, in silico tools can integrate information across multiple scales, from gene expression to crop stands. These tools also allow us to evaluate things that are difficult to measure, such as phenotypes that do not yet exist, or future climates. In silico biology will be an essential tool in 21st Century biology, which will have access to huge amounts of data at multiple scales that can be used to try to understand incredibly complex systems, such as the human brain or roots interacting with living soil. Our main in silico tool is SimRoot, developed over the past 25 years to understand how root phenotypes affect soil resource capture.
Check out a SimRoot model below:
You have been working on breeding plants that have improved yield in soils with low fertility. What have you achieved in this work?
In collaboration with crop breeders and colleagues in various nations we have developed improved common bean lines with better yield under drought and low soil fertility that are being deployed in Africa and Latin America, improved soybean lines with better yield in soils with low phosphorus being deployed in Africa and Asia, and are now working with maize breeders in Africa to develop lines with better yield under drought and low nitrogen stress. Many crop breeders are using our methods for root phenotyping to target root phenotypes in their selection regimes in multiple crops.
What piece of advice do you have for early career researchers?
You are at the forefront of an unprecedented challenge we face as a species – how to sustain 10 billion people in a degrading environment. Plant biologists are an essential part of the effort to reshape how we live on this planet. Do not doubt the importance of your efforts. Do not lose sight of the very real human impact of your scientific choices. Do not be deterred by the gamesmanship and ‘primate politics’ of science. You can make a difference. We need you.