

 Dr. Sarah Schmidt (@BananarootsBlog), Researcher and Science Communicator at The Sainsbury Laboratory Science. Sarah got hooked on both banana research and science writing when she joined a banana Fusarium wilt field trip in Indonesia as a Fusarium expert. She began blogging at https://bananaroots.wordpress.com and just filmed her first science video. She speaks at public events like the Pint of Science and Norwich Science Festival.
Dr. Sarah Schmidt (@BananarootsBlog), Researcher and Science Communicator at The Sainsbury Laboratory Science. Sarah got hooked on both banana research and science writing when she joined a banana Fusarium wilt field trip in Indonesia as a Fusarium expert. She began blogging at https://bananaroots.wordpress.com and just filmed her first science video. She speaks at public events like the Pint of Science and Norwich Science Festival.
Every morning I slice a banana onto my breakfast cereal.
And I am not alone.
Every person in the UK eats, on average, 100 bananas per year.
Bananas are rich in fiber, vitamins, and minerals like potassium and magnesium. Their high carbohydrate and potassium content makes them a favorite snack for professional sports players; the sugar provides energy and the potassium protects the players from muscle fatigue. Every year, around 5000 kg of bananas are consumed by tennis players at Wimbledon.
But bananas are not only delicious snacks and handy energy kicks. For around 100 million people in Sub-Saharan Africa, bananas are staple crops vital for food security. Staple crops are those foods that constitute the dominant portion of a standard diet and supply the major daily calorie intake. In the UK, the staple crop is wheat. We eat wheat-based products for breakfast (toast, cereals), lunch (sandwich), and dinner (pasta, pizza, beer).
In Uganda, bananas are staple crops. Every Ugandan eats 240 kg bananas per year. That is around 7–8 bananas per day. Ugandans do not only eat the sweet dessert banana that we know; in the East African countries such as Kenya, Burundi, Rwanda, and Uganda, the East African Highland banana, called Matooke, is the preferred banana for cooking. Highland bananas are large and starchy, and are harvested green. They can be cooked, fried, boiled, or even brewed into beer, so have very similar uses wheat in the UK.
In West Africa and many Middle and South American countries, another cooking banana, the plantain, is cooked and fried as a staple crop.
In terms of production, the sweet dessert banana we buy in supermarkets is still the most popular. This banana variety is called Cavendish and makes up 47% of the world’s banana production, followed by Highland bananas (24%) and plantains (17%). Last year, I visited Uganda and I managed to combine the top three banana cultivars in one dish: cooked and mashed Matooke, a fried plantain and a local sweet dessert banana!
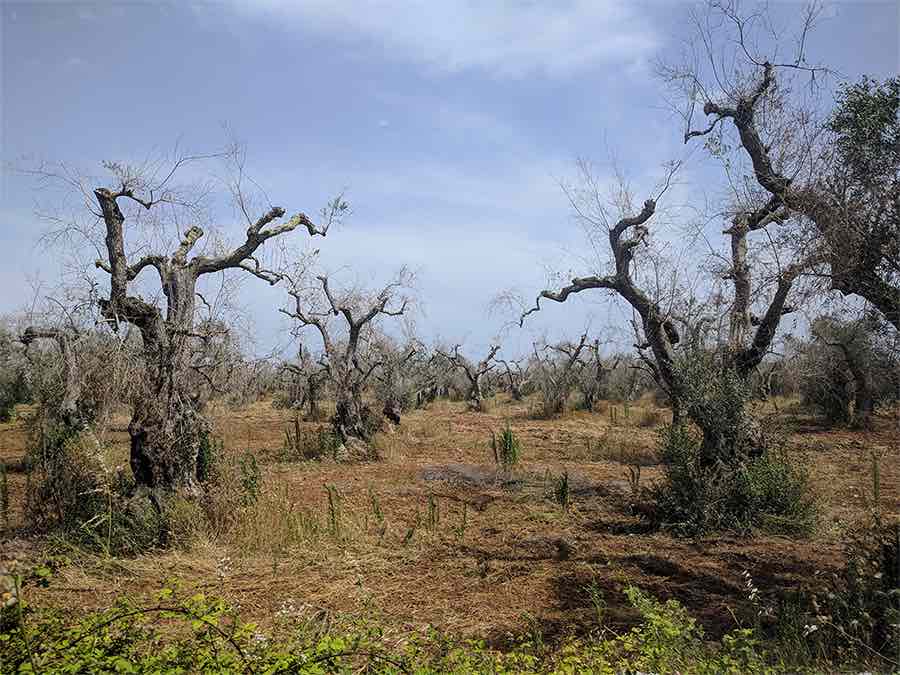
Another important banana cultivar is the sweet dessert banana cultivar Gros Michel, which constitutes 12% of the global production. Gros Michel used to be the most popular banana cultivar worldwide until an epidemic of Fusarium wilt disease devastated the banana export plantations in the so-called “banana republics” in Middle America (Panama, Honduras, Guatemala, Costa Rica) in the 1950s.
Fusarium wilt disease is caused by the soil-borne fungus Fusarium oxysporum f. sp. cubense (FOC). The fungus infects the roots of the banana plants and grows up through the water-conducting, vascular system of the plant. Eventually, this blocks the water transport of the plant and the banana plants start wilting before they can set fruits.
The Fusarium wilt epidemic in Middle America marked the rise of the Cavendish, the only cultivar that could be grown on soils infested with FOC. The fact that they are also the highest yielding banana cultivar quickly made Cavendish the most popular banana variety, both for export and for local consumption.
Currently, Fusarium wilt is once again the biggest threat to worldwide banana production. In the 1990s, a new race of Fusarium wilt – called Tropical Race 4 (TR4) – occurred in Cavendish plantations in Indonesia and Malaysia. Since then, TR4 has spread to the neighboring countries (Taiwan, the Philippines, China, and Australia), but also to distant locations such as Pakistan, Oman, Jordan, and Mozambique.
In Mozambique, the losses incurred by TR4 amounted to USD 7.5 million within just two years. Other countries suffer even more; TR4 causes annual economic losses of around USD 14 million in Malaysia, USD 121 million in Indonesia, and in Taiwan the annual losses amount to a whopping USD 253 million.
TR4 is not only diminishing harvests. It also raises the price of production, because producers have to implement expensive preventative measures and treatments of affected plantations. These preventive measures and treatments are part of the discussion at The World Banana Forum (WBF). The WBF is a permanent platform for all stakeholders of the banana supply chain, and is housed by the United Nation’s Food and Agricultural Organization (FAO). In December 2013, the WBF created a special taskforce to deal with the threat posed by TR4.
Despite its massive impact on banana production, we know very little about the pathogen that is causing Fusarium wilt disease. We don’t know how it spreads, why the new TR4 is so aggressive, or how we can stop it.
Breeding bananas is incredibly tedious, because edible cultivars are sterile and do not produce seeds. I am therefore exploring other ways to engineer resistance in banana against Fusarium wilt. As a scientist in the 2Blades group at The Sainsbury Laboratory, I am investigating how we can transfer resistance genes from other crop species into banana and, more recently, I have been investigating bacteria that are able to inhibit the growth and sporulation of F. oxysporum. These biologicals would be a fast and cost-effective way of preventing or even curing Fusarium wilt disease.
Twitter: @BananarootsBlog
Email: mailto:sarah.schmidt@tsl.ac.uk
Website: https://bananaroots.wordpress.com
In October 2015, researchers from around the world came together in Iguassu Falls, Brazil, for the Stress Resilience Symposium, organized by the Global Plant Council and the Society for Experimental Biology (SEB), to discuss the current research efforts in developing plants resistant to the changing climate. (See our blog by GPC’s Lisa Martin for more on this meeting!)
Building on the success of the meeting, the Global Plant Council team and attendees compiled a set of papers to provide a powerful call to action for stress resilience scientists around the world to come together to tackle some of the biggest challenges we will face in the future. These four papers were published in the Open Access journal Food and Energy Security alongside an editorial about the Global Plant Council.
In the editorial, the Global Plant Council team (Lisa Martin, Sarah Jose, and Ruth Bastow) introduce readers to the Global Plant Council mission, and describe the Stress Resilience initiative, the meeting, and introduce the papers that came from it.
In the first of the commentaries, Matthew Gilliham (University of Adelaide), Scott Chapman (CSIRO), Lisa Martin, Sarah Jose, and Ruth Bastow discuss ‘The case for evidence-based policy to support stress-resilient cropping systems‘, commenting on the important relationships between research and policy and how each must influence the other.
Global Plant Council President Bill Davies (Lancaster University) and CIMMYT‘s Jean-Marcel Ribaut outline the ways in which research can be translated into locally adapted agricultural best practices in their article, ‘Stress resilience in crop plants: strategic thinking to address local food production problems‘.
In the next paper, ‘Harnessing diversity from ecosystems to crops to genes‘, Vicky Buchanan-Wollaston (University of Warwick), Zoe Wilson (University of Nottingham), François Tardieu (INRA), Jim Beynon (University of Warwick), and Katherine Denby (University of York) describe the challenges that must be overcome to promote effective and efficient international research collaboration to develop new solutions and stress resilience plants to enhance food security in the future.
University of Queensland‘s Andrew Borrell and CIMMYT‘s Matthew Reynolds discuss how best to bring together researchers from different disciplines, highlighting great examples of this in their paper, ‘Integrating islands of knowledge for greater synergy and efficiency in crop research‘.
In all of these papers, the authors suggest practical short- and long-term action steps and highlight ways in which the Global Plant Council could help to bring researchers together to coordinate these changes most effectively.
Read the papers in Food and Energy Security here.
Another fantastic year of discovery is over – read on for our 2016 plant science top picks!

A Zostera marina meadow in the Archipelago Sea, southwest Finland. Image credit: Christoffer Boström (Olsen et al., 2016. Nature).
The year began with the publication of the fascinating eelgrass (Zostera marina) genome by an international team of researchers. This marine monocot descended from land-dwelling ancestors, but went through a dramatic adaptation to life in the ocean, in what the lead author Professor Jeanine Olsen described as, “arguably the most extreme adaptation a terrestrial… species can undergo”.
One of the most interesting revelations was that eelgrass cannot make stomatal pores because it has completely lost the genes responsible for regulating their development. It also ditched genes involved in perceiving UV light, which does not penetrate well through its deep water habitat.
Read the paper in Nature: The genome of the seagrass Zostera marina reveals angiosperm adaptation to the sea.
BLOG: You can find out more about the secrets of seagrass in our blog post.
Plants are known to form new organs throughout their lifecycle, but it was not previously clear how they organized their cell development to form the right shapes. In February, researchers in Germany used an exciting new type of high-resolution fluorescence microscope to observe every individual cell in a developing lateral root, following the complex arrangement of their cell division over time.
Using this new four-dimensional cell lineage map of lateral root development in combination with computer modelling, the team revealed that, while the contribution of each cell is not pre-determined, the cells self-organize to regulate the overall development of the root in a predictable manner.
Watch the mesmerizing cell division in lateral root development in the video below, which accompanied the paper:
Read the paper in Current Biology: Rules and self-organizing properties of post-embryonic plant organ cell division patterns.
In March, a Spanish team of researchers revealed how the anti-wilting molecular machinery involved in preserving cell turgor assembles in response to drought. They found that a family of small proteins, the CARs, act in clusters to guide proteins to the cell membrane, in what author Dr. Pedro Luis Rodriguez described as “a kind of landing strip, acting as molecular antennas that call out to other proteins as and when necessary to orchestrate the required cellular response”.
Read the paper in PNAS: Calcium-dependent oligomerization of CAR proteins at cell membrane modulates ABA signaling.
*If you’d like to read more about stress resilience in plants, check out the meeting report from the Stress Resilience Forum run by the GPC in coalition with the Society for Experimental Biology.*
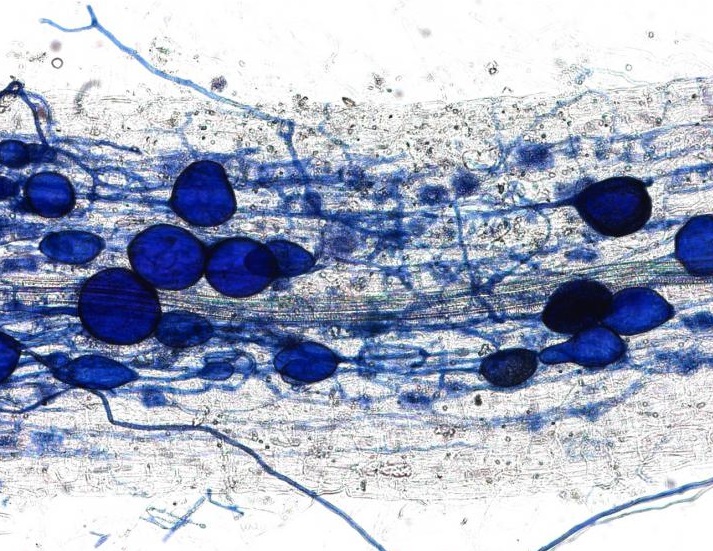
This plant root is infected with arbuscular mycorrhizal fungi. Image credit: University of Zurich.
In April, we received an amazing insight into the ‘decision-making ability’ of plants when a Swiss team discovered that plants can punish mutualist fungi that try to cheat them. In a clever experiment, the researchers provided a plant with two mutualistic partners; a ‘generous’ fungus that provides the plant with a lot of phosphates in return for carbohydrates, and a ‘meaner’ fungus that attempts to reduce the amount of phosphate it ‘pays’. They revealed that the plants can starve the meaner fungus, providing fewer carbohydrates until it pays its phosphate bill.
Author Professor Andres Wiemsken explains: “The plant exploits the competitive situation of the two fungi in a targeted manner, triggering what is essentially a market-based process determined by cost and performance”.
Read the paper in Ecology Letters: Options of partners improve carbon for phosphorus trade in the arbuscular mycorrhizal mutualism.
The transition of ancient plants from water onto land was one of the most important events in our planet’s evolution, but required a massive change in plant biology. Suddenly plants risked drying out, so had to develop new ways to survive drought.
In May, an international team discovered a key gene in moss (Physcomitrella patens) that allows it to tolerate dehydration. This gene, ANR, was an ancient adaptation of an algal gene that allowed the early plants to respond to the drought-signaling hormone ABA. Its evolution is still a mystery, though, as author Dr. Sean Stevenson explains: “What’s interesting is that aquatic algae can’t respond to ABA: the next challenge is to discover how this hormone signaling process arose.”
Read the paper in The Plant Cell: Genetic analysis of Physcomitrella patens identifies ABSCISIC ACID NON-RESPONSIVE, a regulator of ABA responses unique to basal land plants and required for desiccation tolerance.
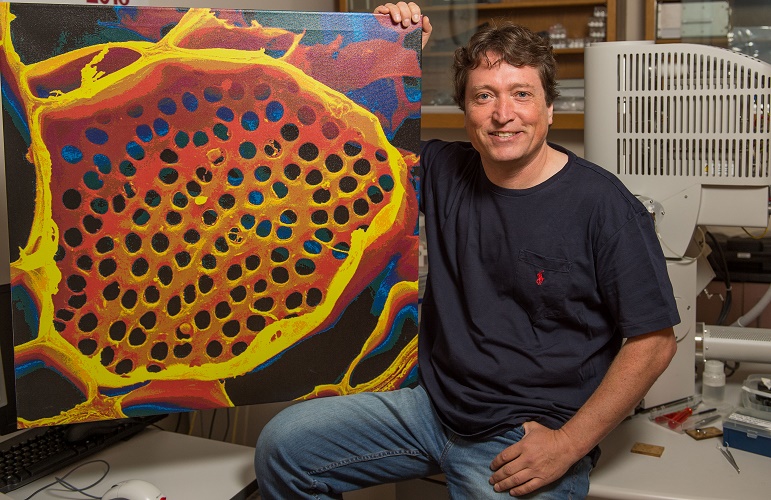
Professor Michael Knoblauch shows off a microscope image of phloem tubes. Image credit: Washington State University.
Sometimes revisiting old ideas can pay off, as a US team revealed in June. In 1930, Ernst Münch hypothesized that transport through the phloem sieve tubes in the plant vascular tissue is driven by pressure gradients, but no-one really knew how this would account for the massive pressure required to move nutrients through something as large as a tree.
Professor Michael Knoblauch and colleagues spent decades devising new methods to investigate pressures and flow within phloem without disrupting the system. He eventually developed a suite of techniques, including a picogauge with the help of his son, Jan, to measure tiny pressure differences in the plants. They found that plants can alter the shape of their phloem vessels to change the pressure within them, allowing them to transport sugars over varying distances, which provided strong support for Münch flow.
Read the paper in eLife: Testing the Münch hypothesis of long distance phloem transport in plants.
BLOG: We featured similar work (including an amazing video of the wound response in sieve tubes) by Knoblauch’s collaborator, Dr. Winfried Peters, on the blog – read it here!
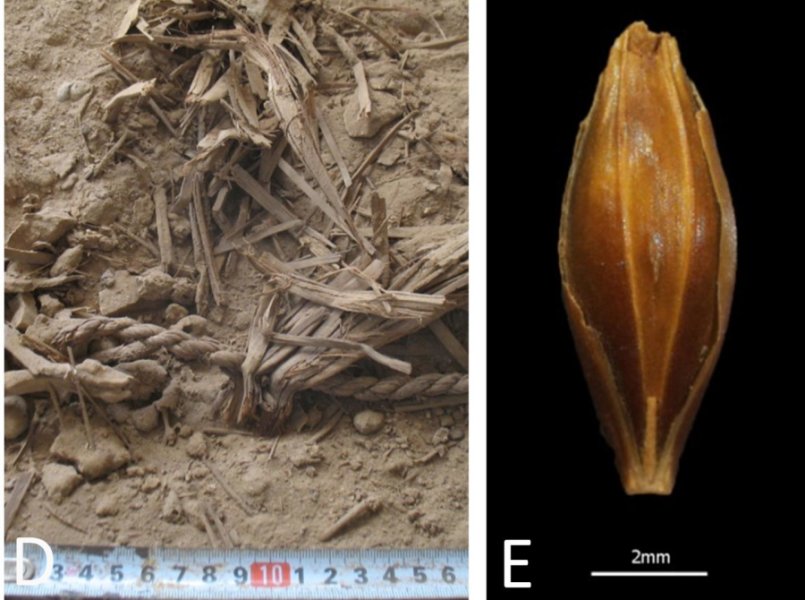

Preserved remains of rope, seeds, reeds and pellets (left), and a desiccated barley grain (right) found at Yoram Cave in the Judean Desert. Credit: Uri Davidovich and Ehud Weiss.
In July, an international and highly multidisciplinary team published the genome of 6,000-year-old barley grains excavated from a cave in Israel, the oldest plant genome reconstructed to date. The grains were visually and genetically very similar to modern barley, showing that this crop was domesticated very early on in our agricultural history. With more analysis ongoing, author Dr. Verena Schünemann predicts that “DNA-analysis of archaeological remains of prehistoric plants will provide us with novel insights into the origin, domestication and spread of crop plants”.
Read the paper in Nature Genetics: Genomic analysis of 6,000-year-old cultivated grain illuminates the domestication history of barley.
BLOG: We interviewed Dr. Nils Stein about this fascinating work on the blog – click here to read more!
Another exciting cereal paper was published in August, when an Australian team revealed that C4 photosynthesis occurs in wheat seeds. Like many important crops, wheat leaves perform C3 photosynthesis, which is a less efficient process, so many researchers are attempting to engineer the complex C4 photosynthesis pathway into C3 crops.
This discovery was completely unexpected, as throughout its evolution wheat has been a C3 plant. Author Professor Robert Henry suggested: “One theory is that as [atmospheric] carbon dioxide began to decline, [wheat’s] seeds evolved a C4 pathway to capture more sunlight to convert to energy.”
Read the paper in Scientific Reports: New evidence for grain specific C4 photosynthesis in wheat.
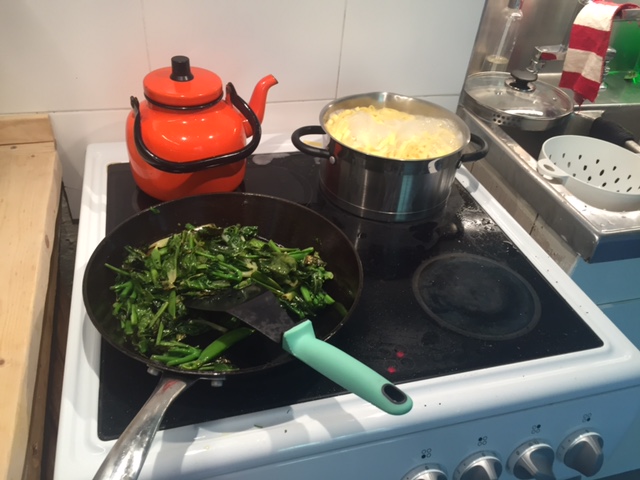

Professor Stefan Jansson cooks up “Tagliatelle with CRISPRy fried vegetables”. Image credit: Stefan Jansson.
September marked an historic event. Professor Stefan Jansson cooked up the world’s first CRISPR meal, tagliatelle with CRISPRy fried vegetables (genome-edited cabbage). Jansson has paved the way for CRISPR in Europe; while the EU is yet to make a decision about how CRISPR-edited plants will be regulated, Jansson successfully convinced the Swedish Board of Agriculture to rule that plants edited in a manner that could have been achieved by traditional breeding (i.e. the deletion or minor mutation of a gene, but not the insertion of a gene from another species) cannot be treated as a GMO.
Read more in the Umeå University press release: Umeå researcher served a world first (?) CRISPR meal.
BLOG: We interviewed Professor Stefan Jansson about his prominent role in the CRISPR/GM debate earlier in 2016 – check out his post here.
*You may also be interested in the upcoming meeting, ‘New Breeding Technologies in the Plant Sciences’, which will be held at the University of Gothenburg, Sweden, on 7-8 July 2017. The workshop has been organized by Professor Jansson, along with the GPC’s Executive Director Ruth Bastow and Professor Barry Pogson (Australian National University/GPC Chair). For more info, click here.*
Phytochromes help plants detect day length by sensing differences in red and far-red light, but a UK-Germany research collaboration revealed that these receptors switch roles at night to become thermometers, helping plants to respond to seasonal changes in temperature.
Dr Philip Wigge explains: “Just as mercury rises in a thermometer, the rate at which phytochromes revert to their inactive state during the night is a direct measure of temperature. The lower the temperature, the slower phytochromes revert to inactivity, so the molecules spend more time in their active, growth-suppressing state. This is why plants are slower to grow in winter”.
Read the paper in Science: Phytochromes function as thermosensors in Arabidopsis.
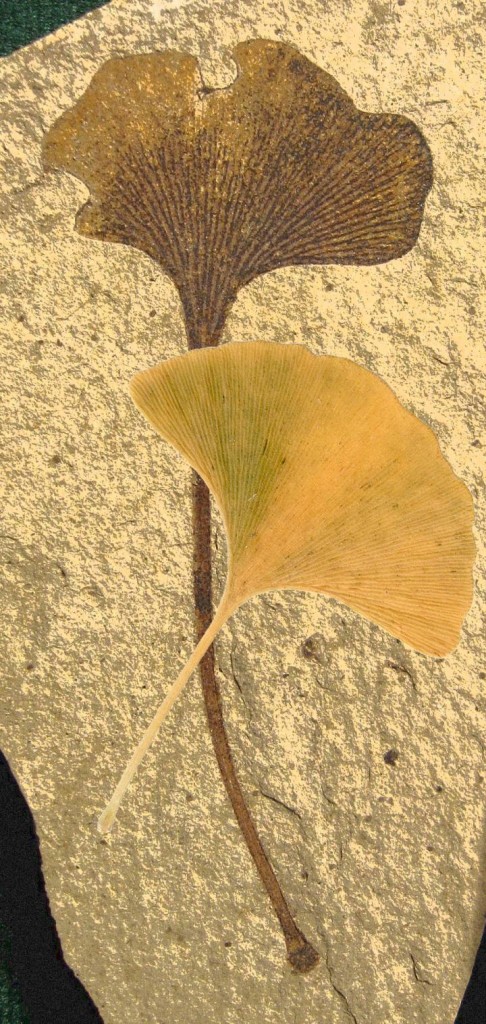

A fossil ginkgo (Ginkgo biloba) leaf with its modern counterpart. Image credit: Gigascience.
In November, a Chinese team published the genome of Ginkgo biloba¸ the oldest extant tree species. Its large (10.6 Gb) genome has previously impeded our understanding of this living fossil, but researchers will now be able to investigate its ~42,000 genes to understand its interesting characteristics, such as resistance to stress and dioecious reproduction, and how it remained almost unchanged in the 270 million years it has existed.
Author Professor Yunpeng Zhao said, “Such a genome fills a major phylogenetic gap of land plants, and provides key genetic resources to address evolutionary questions [such as the] phylogenetic relationships of gymnosperm lineages, [and the] evolution of genome and genes in land plants”.
Read the paper in GigaScience: Draft genome of the living fossil Ginkgo biloba.
The year ended with another fascinating discovery from a Danish team, who used fluorescent tags and microscopy to confirm the existence of metabolons, clusters of metabolic enzymes that have never been detected in cells before. These metabolons can assemble rapidly in response to a stimulus, working as a metabolic production line to efficiently produce the required compounds. Scientists have been looking for metabolons for 40 years, and this discovery could be crucial for improving our ability to harness the production power of plants.
Read the paper in Science: Characterization of a dynamic metabolon producing the defense compound dhurrin in sorghum.
Another amazing year of science! We’re looking forward to seeing what 2017 will bring!
P.S. Check out 2015 Plant Science Round Up to see last year’s top picks!
This week’s blog post was written by Dr Damiano Martignago, a genome editing specialist at Rothamsted Research.
Genome editing technologies comprise a diverse set of molecular tools that allow the targeted modification of a DNA sequence within a genome. Unlike “traditional” breeding, genome editing does not rely on random DNA recombination; instead it allows the precise targeting of specific DNA sequences of interest. Genome editing approaches induce a double strand break (DSB) of the DNA molecule at specific sites, activating the cell’s DNA repair system. This process could be either error-prone, thus used by scientists to deactivate “undesired” genes, or error-free, enabling target DNA sequences to be “re-written” or the insertion of DNA fragments in a specific genomic position.
The most promising among the genome editing technologies, CRISPR/Cas9, was chosen as Science’s 2015 Breakthrough of the Year. Cas9 is an enzyme able to target a specific position of a genome thanks to a small RNA molecule called guide RNA (gRNA). gRNAs are easy to design and can be delivered to cells along with the gene encoding Cas9, or as a pre-assembled Cas9-gRNA protein-RNA complex. Once inside the cell, Cas9 cuts the target DNA sequence homologous to the gRNAs, producing DSBs.
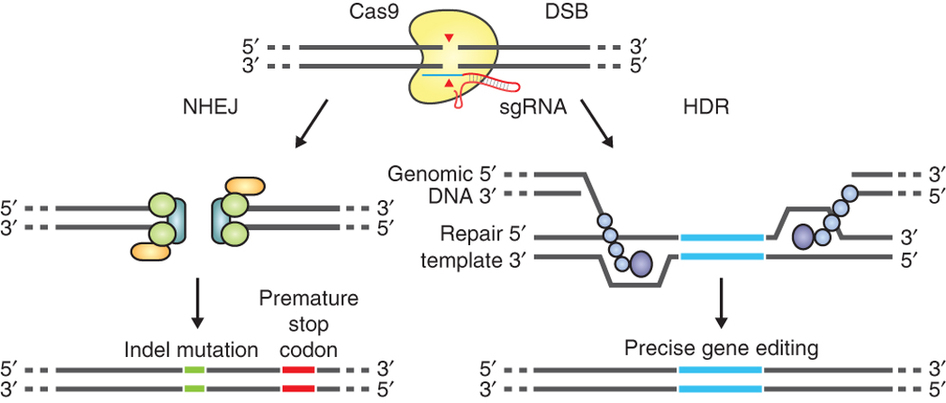
The guide RNA (sgRNA) directs Cas9 to a specific region of the genome, where it induces a double-strand break in the DNA. On the left, the break is repaired by non-homologous-end joining, which can result in insertion/deletion (indel) mutations. On the right, the homologous-directed recombination pathway creates precise changes using a supplied template DNA. Credit: Ran et al. (2013). Nature Protocols.
Together with the increased data availability on crop genomes, genome editing techniques such as CRISPR are allowing scientists to carry out ambitious research on crop plants directly, building on the knowledge obtained during decades of investigation in model plants.
The concept of CRISPR was first tested in crops by generating cultivars that are resistant to herbicides, as this is an easy trait to screen for and identify. One of the first genome-edited crops, a herbicide-resistant oilseed rape produced by Cibus, has already been grown and harvested in the USA in 2015.

Researchers used CRISPR to engineer a wheat variety resistant to powdery mildew (shown here), a major disease of this crop. Image credit: NY State IPM Program. Used under license: CC BY 2.0.
Using CRISPR, scientists from the Chinese Academy of Sciences produced a wheat variety resistant to powdery mildew, one of the major diseases in wheat. Similarly, another Chinese research group exploited CRISPR to produce a rice line with enhanced rice blast resistance that will help to reduce the amount of fungicides used in rice farming. CRISPR/Cas9 has also been already applied to maize, tomato, potato, orange, lettuce, soybean and other legumes.
Genome editing could also revolutionize the management of viral plant disease. The CRISPR/Cas9 system was originally discovered in bacteria, where it provided them with molecular immunity against viruses, but it can also be moved into plants. Scientists can transform plants to produce the Cas9 and gRNAs that target viral DNA, reducing virus accumulation; alternatively, they can suppress those plant genes that are hijacked by the virus to mediate its own diffusion in the plants. Since most plants are defenseless against viruses and there are no chemical controls available for plant viruses, the main method to stop the spread of these diseases is still the destruction of the infected plant. For the first time in history, scientists have an effective weapon to fight back against plant viruses.
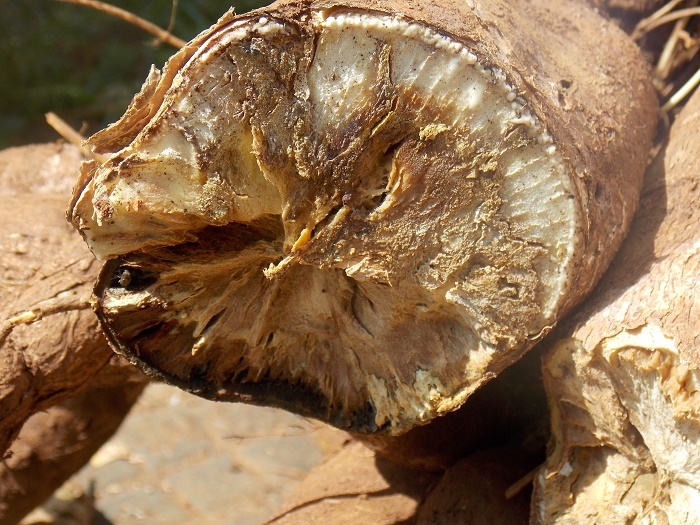

The cassava brown streak disease virus can destroy cassava crops, threatening the food security of the 300 million people who rely on this crop in Africa. Image credit: Katie Tomlinson (for more on this topic, read her blog here).
Genome editing will be particularly useful in the genetic improvement of many crops that are propagated mainly by vegetative reproduction, and so very difficult to improve by traditional breeding methods involving crossing (e.g. cassava, banana, grape, potato). For example, using TALENs, scientists from Cellectis edited a potato line to minimize the accumulation of reducing sugars that may be converted into acrylamide (a possible carcinogen) during cooking.
One of the hypothesized risks of using CRISPR/Cas9 is the potential targeting of undesired DNA regions, called off-targets. It is possible to limit the potential for off-targets by designing very specific gRNAs, and all of the work published so far either did not detect any off-targets or, if detected, they occurred at a very low frequency. The number of off-target mutations produced by CRISPR/Cas9 is therefore minimal, especially if compared with the widely accepted random mutagenesis of crops used in plant breeding since the 1950s.
Genome editing is interesting from a regulatory point of view too. After obtaining the desired heritable mutation using CRISPR/Cas9, it is possible to remove the CRISPR/Cas9 integrated vectors from the genome using simple genetic segregation, leaving no trace of the genome modification other than the mutation itself. This means that some countries (including the USA, Canada, and Argentina) consider the products of genome editing on a case-by-case basis, ruling that a crop is non-GM when it contains gene combinations that could have been obtained through crossing or random mutation. Many other countries are yet to issue an official statement on CRISPR, however.
Recently, scientists showed that is possible to edit the genome of plants without adding any foreign DNA and without the need for bacteria- or virus-mediated plant transformation. Instead, a pre-assembled Cas9-gRNA ribonucleoprotein (RNP) is delivered to plant cells in vitro, which can edit the desired region of the genome before being rapidly degraded by the plant endogenous proteases and nucleases. This non-GM approach can also reduce the potential of off-target editing, because of the minimal time that the RNP is present inside the cell before being degraded. RNP-based genome editing has been already applied to tobacco plants, rice, and lettuce, as well as very recently to maize.
In conclusion, genome editing techniques, and CRISPR/Cas9 in particular, offers scientists and plant breeders a flexible and relatively easy approach to accelerate breeding practices in a wide variety of crop species, providing another tool that we can use to improve food security in the future.
For more on CRISPR, check out this recent TED Talk from Ellen Jorgensen:
Dr Damiano Martignago is a plant molecular biologist who graduated from Padua University, Italy, with a degree in Food Biotechnology in 2009. He obtained his PhD in Biology at Roma Tre University in 2014. His experience with CRISPR/Cas9 began in the lab of Prof. Fabio Fornara (University of Milan), where he used CRISPR/Cas9 to target photoperiod genes of interest in rice and generate mutants that were not previously available. He recently moved to Rothamsted Research, UK, where he works as Genome Editing Specialist, transferring CRISPR/Cas9 technology to hexaploid bread wheat with the aim of improving the efficiency of genome editing in this crop. He is actively involved with AIRIcerca (International Association of Italian Scientists), disseminating and promoting scientific news.
This week we speak to Tim Williams, the Business Manager of Farming Futures and Research Fund Development Manager at Aberystwyth University, UK.
Could you give a brief introduction to Farming Futures and its mission?
Farming Futures is an independent, UK-based, inclusive agri-food supply chains alliance. Our mission is to work with researchers and industry to share knowledge, with the aim of improving the sustainability and productive efficiency of agriculture, all within the context of healthy, high-quality food.
What is the history of the organization?
Farming Futures started with an idea by Professor Wayne Powell in 2009 (then the director of the Institute of Biological, Environmental and Rural Sciences (IBERS) at Aberystwyth) in discussion with Mark Price, who was the Managing Director of British supermarket chain Waitrose. It was launched in 2010, starting out as the Centre of Excellence for UK Farming (CEUKF). Waitrose seed-funded Farming Futures, and since then we have received support from the Agriculture and Horticulture Development Board (AHDB) and Innovate UK.
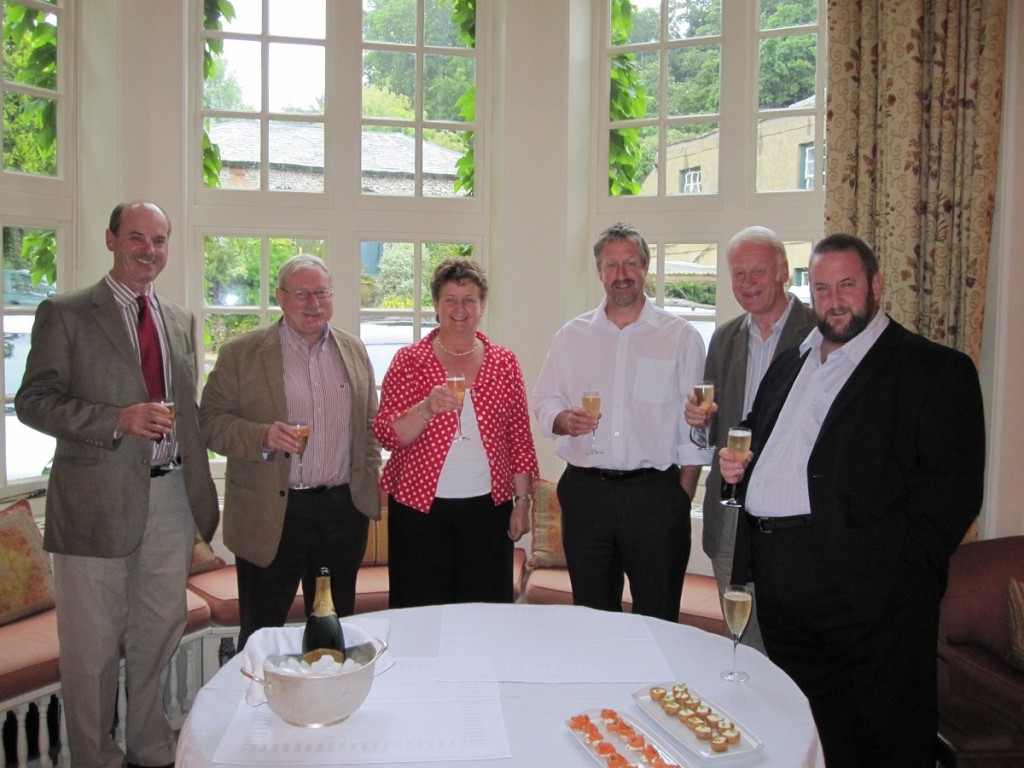

The inauguration meeting of Farming Futures in 2009, then known as the Centre of Excellence for UK Farming. Left-Right: Tim Williams, Wayne Powell, Heather Jenkins, David Davies, Philip Morgan, Jamie Newbold.
How has plant and crop research been integrated into the recommendations presented by Farming Futures?
Plant science is the fundamental driver for agri-food development. We work closely with industry, as well as the AHDB and other farm advisory bodies across the UK to inform them about new developments. Accelerated, directed breeding programs using genomic and phenomic technologies are helping us to develop new varieties that offer more productive, more resilient, environmentally friendly plants – not just as food crops, but also for soil quality, nutrient retention, flood reduction, energy biomass, renewable chemistry, and a host of other desirable characteristics.
Historically, to paraphrase a fellow botanist, we have bred ‘needy, greedy plants’ that deplete resources and need lots of nasty chemicals to keep them growing. Now scientists are mining the genomes of crop ancestors to rediscover the genetic traits we unwittingly threw away on the route to increased yield.
What roles do research partners such as universities play?
We work together in a pre-competitive way to enable research, and to represent farming within agri-food policy – researchers from different organizations can collaborate thanks to our partners’ trusting relationships with each other. Collaborations in science are vital because the problems our global society faces are multi-factorial, non-linear and multi-disciplinary. They are far too complex for the typical university research team, working alone, to address efficiently. We need the equivalent of the CERN Large Hadron Collider project for agri-food.
In addition to helping researchers to bring in millions of pounds worth of applied research projects (at least £12 million, but it is notoriously difficult to find out what industry is funding), Farming Futures helped to establish the government-funded Agri-Food Tech Centres of Innovation for a total of around £90 million, bringing in industry to co-fund and support three of the four: the Agrimetrics Centre, Agri-Epi-Centre and Centre of Innovation Excellence in Livestock. In time, these Centres will catalyze a lot of collaborative research and will help stimulate innovation and technology uptake by industry.
…Economic returns on R&D are about 27 X investment but takes an average of 23 yrs for R&D innovation to be taken up by agriculture. 2/2
— DefraChiefScientist (@DefraChiefScien) January 27, 2016
What climate change challenges will farmers face? Are there any specific challenges that Farming Futures can address?
Farming Futures and its network brings together scientists from different disciplines to discuss these problems and potential solutions. For instance, people from the UK’s national weather service (the Met Office) and some of the biggest food retailers and processors in the world come together at our conferences and workshops to think through scenarios and solutions. These solutions include breeding crops for increased resilience, not just peak yield. We are running out of fungicides that work efficiently, in the same way that we are running out of antibiotics; however, some very clever scientists have worked out some potential solutions that are more environmentally sound, so I am an optimist.
This problem solving is best done at the supply-chain level as it brings in a wider expertise. As I repeat often, a colleague once said to the board of one of the world’s biggest brewers, “No barley = no beer = no business”, inferring the question, “What are you doing to ensure that barley growers are going to be able to supply you in the future?”
Your website has an interesting study from 2011 highlighting six potential jobs of the future, including geoengineer, energy farming, web 3.0 farm host, pharmer, etc. How can students direct their skill development to meet the needs of the future?
There are many emerging jobs and skills, but each of these named jobs from 2011 are actually in practice now. The web 3.0 has now become web 4.0, which is the “internet of things”, with data collection from lots of devices including drones for precision agriculture and robots for weeding and picking crops.
The future of agri-food is in big data, including consumer behavior, weather forecasting, genomics, phenomics, and real-time analysis of the growth progress of plants and animals on-farm. We need more electronic and mechanical engineers with an understanding of biology, as well as more biologists who work within the agri-food industries and in government policy development.
What are you currently working on?
We are currently working with partners on a number of projects across the Agri-Food Tech Centres and trying to form more research collaborations. One of our big projects is The National Library for Agri-Food. I am currently working with web developers and experts from Jisc and the British Library to scope the requirements and to build a demonstration web site.
Finally, I would just like to add that we are open to collaborations across agri-food supply chains and will work to foster them, either openly or privately as appropriate.
In addition to IBERS, Farming Futures has four founding members (Northern Ireland’s Agri-Food and Biosciences Institute (AFBI), Harper Adams University (HAU), NIAB with East Malling Research (NIAB-EMR), and Scotland’s Rural College (SRUC)) and an influential Steering Board, chaired by Lord Curry of Kirkharle, who is very well known and respected in UK government and farming.
This week’s post was written by Katie Tomlinson, a PhD student at the University of Bristol, UK, who spent three months as an intern at the National Crops Resource Research Institute in Uganda. She fills us in on the important research underway at the Institute, and how they communicate their important results to local farmers and benefit rural communities.
Over the summer, I had a great time at the National Crops Resources Research Institute (NaCRRI) in Uganda. I’m currently in the second year of my PhD at the University of Bristol, UK, where I’m researching how the cassava brown streak disease (CBSD) viruses are able to cause symptoms, replicate and move inside plants. I was lucky enough to be given a placement at NaCRRI as part of the South West Doctoral Training Partnership Professional Internship for PhD Students (PIPS) scheme, to experience the problem for myself, see the disease in the field, meet the farmers affected and investigate the possible solutions.
Cassava is a staple food crop for approximately 300 million people in Africa. It is resilient to seasonal drought, can be grown on poor soils and harvested when needed. However, cassava production is seriously threatened by CBSD, which causes yellow patches (chlorosis) to form on leaves and areas of tubers to die (necrosis), rot and become inedible.
Despite being identified in coastal Tanzania 80 years ago, CBSD has only been a serious problem for Uganda in the last 10 years, where it was the most important crop disease in 2014–2015. The disease has since spread across East Africa and threatens the food security of millions of people.
NaCRRI is a government institute, which carries out research to protect and improve the production of key crops, including cassava. The focus is on involving farmers in this process so that the best possible crop varieties and practices are available to them. Communication between researchers and farmers is therefore vital, and it was this that I wanted to assist with.
When I arrived I was welcomed warmly into the root crop team by the team leader Dr Titus Alicai, who came up with a whole series of activities to give me a real insight into CBSD. I was invited to field sites across Uganda, where I got to see CBSD symptoms in the flesh! I helped to collect data for the 5CP project, which is screening different cassava varieties from five East and Southern African countries for CBSD and cassava mosaic disease (CMD) resistance. I helped to score plants for symptoms and was fascinated by the variability of disease severity in different varieties. The main insight I gained is that the situation is both complex and dynamic, with some plants appearing to be disease-free while others were heavily infected. There are also different viral strains found across different areas, and viral populations are also continually adapting. The symptoms also depend on environmental conditions, which are unpredictable.
I also got to see super-abundant whiteflies, which transmit viruses, and understand how their populations are affected by environmental conditions. These vectors are also complex; they are expanding into new areas and responding to changing environmental conditions.
It has been fascinating to learn how NaCRRI is tackling the CBSD problem through screening different varieties in the 5CP project, breeding new varieties in the NEXTGEN cassava project, providing clean planting material and developing GM cassava.
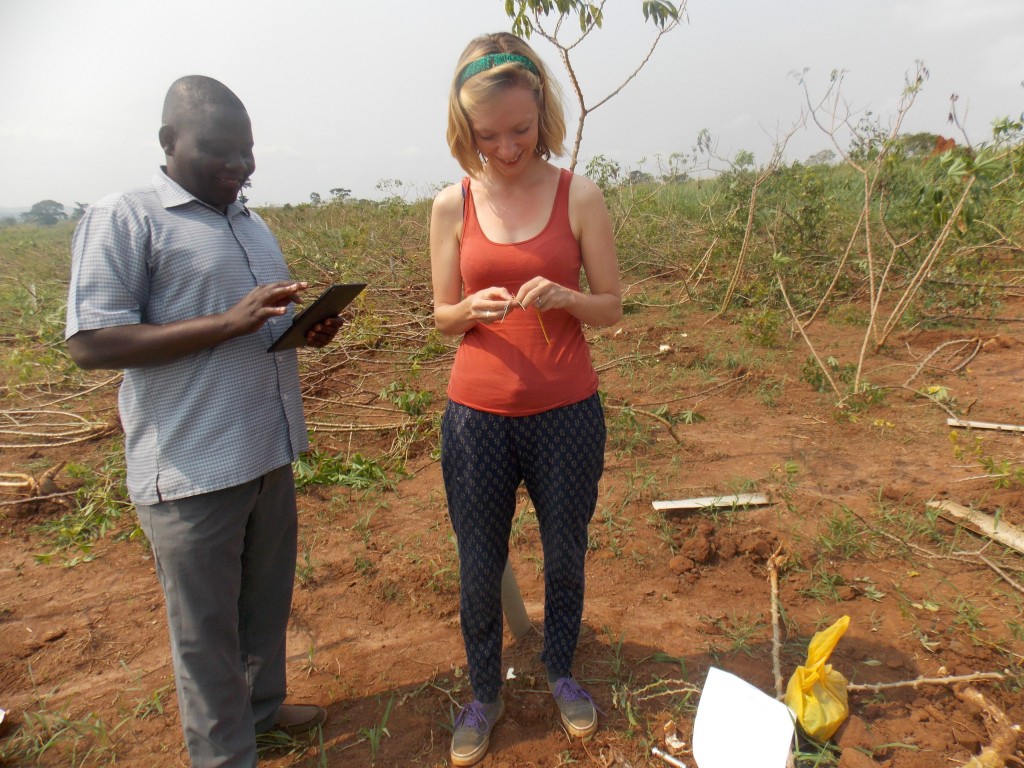

Tagging cassava plants free from Cassava brown streak disease for breeding. Image credit: Katie Tomlinson.
In each of these projects, communication with local farmers is crucial. I’ve had the opportunity to meet farmers directly affected, some of whom have all but given up on growing cassava.
Communicating has not been easy, as there are over 40 local languages. I had to adapt and learn from those around me. For example, in the UK we have a habit of emailing everything, whereas in Uganda I had to talk to people to hear about what was going on. This is all part of the experience and something I’ll definitely be brining back to the UK! I’ve had some funny moments too… during harvesting the Ugandans couldn’t believe how weak I was; I couldn’t even cut one cassava open!
I’m going to treasure my experiences at NaCRRI. The insights into CBSD are already helping me to plan experiments, with more real-world applications. I can now see how all the different elements (plant–virus–vector–environment–human) interact, which is something you can’t learn from reading papers alone!
Working with the NaCRRI team has given me the desire and confidence to collaborate with an international team. I’ve formed some very strong connections and hope to have discussions about CBSD with them throughout my PhD and beyond. It’s really helped to strengthen collaborations between our lab work in Bristol and researchers working in the field on the disease frontline. This will help our research to be relevant to the current situation and what is happening in the field.
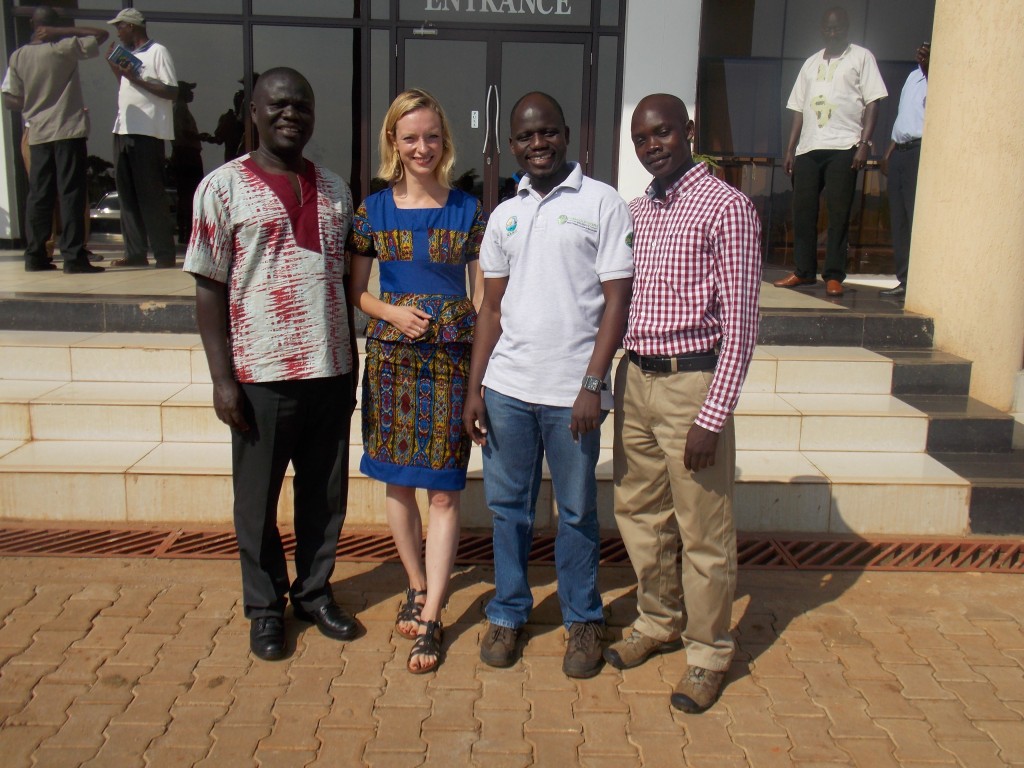

Saying goodbye to new friends: Dr. Titus Alicai (NaCRRI root crops team leader), Phillip Abidrabo (CBSD MSc student) and Dr. Esuma Williams (cassava breeder). Image credit: Katie Tomlinson.
Promoting and supporting plant health will be an important part of how we achieve the United Nations’ Sustainable Development Goals (SDGs). Andrea Powell, Chief Information Officer of the Centre for Agriculture and Biosciences International (CABI) looks at how the CABI-led Plantwise programme is helping to make a difference.
By Andrea Powell
On 26th and 27th July 2016, CABI held its 19th Review Conference. This important milestone in the CABI calendar saw our 48 member countries come together to agree a new medium-term strategy. As always, plant health was a key focus to our discussions, cutting across many of CABI’s objectives. For CABI, with 100 years of experience working in plant health, it has become one of our most important issues, upon which our flagship food security program, Plantwise, has been built.
Plant health can, quite simply, change the lives and livelihoods of millions of people living in rural communities, like smallholder farmers. Human and animal health make headlines, while plant health often falls under the radar, yet, it is crucial to tackling serious global challenges like food security. Promoting and supporting plant health will be an important way to achieve the Sustainable Development Goals (SDGs).
Take, for example, SDG 1, which calls for ‘no poverty’. The UN states that one in five people in developing regions still lives on less than $1.25 a day. We know that many of these people are smallholder farmers. By breaking down the barriers to accessing plant health knowledge, millions of people in rural communities can learn how to grow produce to sell to profitable domestic, regional and international markets.
 SDG 2 focuses on achieving ‘zero hunger’. Almost one billion people go hungry and are left malnourished every day – and many are children. Subsistence farmers, who grow food for their families to eat, can be left with nothing when their crops fail. Access to plant health knowledge can help prevent devastating crop losses and put food on the table.
SDG 2 focuses on achieving ‘zero hunger’. Almost one billion people go hungry and are left malnourished every day – and many are children. Subsistence farmers, who grow food for their families to eat, can be left with nothing when their crops fail. Access to plant health knowledge can help prevent devastating crop losses and put food on the table.
Interestingly, SDG 17 considers ‘partnerships for the goals’ and is critical to the way in which we can harness and share plant health knowledge more widely to help address issues like hunger and poverty. By themselves, individual organizations cannot easily resolve the complicated and interconnected challenges the world faces today. This is why partnership is at the heart of CABI’s flagship plant health programme: Plantwise.
Since its launch in 2011, the goal of Plantwise has been to deliver plant health knowledge to smallholder farmers, ensuring they lose less of what they grow. This, in turn, provides food for their families and improves living conditions in rural communities. Plantwise provides support to governments, helping to make national plant health systems more effective for the farmers who depend on them. Already, Plantwise has reached nearly five million farmers. With additional funding, and by developing new partnerships, we aim to bring relevant plant health information to 30 million farmers by 2020, safeguarding food security for generations to come.
Plantwise ‘plant clinics’ are an important part of the fight against crop losses. Established in much the same way as clinics for human health, farmers visit the clinics with samples of their sick crops. Plant doctors diagnose the problem, making science-based recommendations on ways to manage it. The clinics are owned and operated by over 200 national partner organizations in over 30 countries. At the end of 2015, nearly five thousand plant doctors had been trained.
The Plantwise Knowledge Bank reinforces the plant clinics. Available in over 80 languages, it is an online and offline gateway to plant health information, providing the plant doctors with actionable information. It also collects data about the farmers, their crops and plant health problems. This enables in-country partner organizations to monitor the quality of plant doctor recommendations; to identify new plant health problems – often emerging due to trade or climate change issues; and develop new best-practice guidelines for managing crop losses.

The first ever e-plant clinic, held in Embu Market, Kenya. Credit: Plantwise
The Plantwise flow of information improves knowledge and helps the users involved: farmers can receive crop management advice, and researchers and governments can access data from the field. With a new strategy for 2017–19 agreed, CABI will continue to focus on building strong plant health systems. We are certain that plant health is of central importance to achieving the SDGs and, together in partnership, we look forward to growing the Plantwise program and making a concrete difference to the lives of smallholder farmers.
“A few years ago, I would make ZMW 5000 per year. Last year I got 15 000. I have never missed any plant clinic session. I’ve been very committed, very faithful, because I have seen the benefits.”––Kenny Mwansa, Farmer, Rufunsa District, Zambia.
Take a look at Plantwise in action in Zambia (YouTube):
Meet Linda, a Zambian plant doctor
Learn more about Plantwise at www.plantwise.org.
This week on the blog, Professor Sophien Kamoun describes his work on plant–pathogen interactions at The Sainsbury Lab, UK, and discusses the future of plant disease.
Could you begin by describing the focus of your research on plant pathogens?
We study several aspects of plant–pathogen interactions, ranging from genome-level analyses to mechanistic investigations focused on individual proteins. Our projects are driven by some of the major questions in the field: how do plant pathogens evolve? How do they adapt and specialize to their hosts? How do plant pathogen effectors co-opt host processes?
One personal aim is to narrow the gap between research on the mechanisms and evolution of these processes. We hope to demonstrate how mechanistic research benefits from a robust phylogenetic framework to test specific hypotheses about how evolution has shaped molecular mechanisms of pathogenicity and immunity.

Sudden oak death is caused by the oomycete Phytophthora ramorum. Image from Nichols, 2014. PLOS Biology.
Tree diseases such as sudden oak death, ash dieback and olive quick decline syndrome have been making the news a lot recently. Are diseases like these becoming more common, and if so, why?
It’s well documented that the scale and frequency of emerging plant diseases has increased. There are many factors to blame. Increased global trade is one. Climate change is another. There is no question that we need to increase our surveillance and diagnostics efforts. We’re nowhere near having coordinated responses to new disease outbreaks in plant pathology, especially when it comes to deploying the latest genomics methods. We really need to remedy this.
The wheat blast fungus recently hit Bangladesh. Could you briefly outline how it is being tackled by plant pathologists?
Wheat blast has just emerged this last February in Bangladesh – its first report in Asia. It could spread to neighboring countries and become a major threat to wheat production in South Asia. Thus, we had to act fast. We used an Open Science approach to mobilize collaborators in Bangladesh and the wider blast fungus community, and managed to identify the pathogen strain in just a few weeks. It turned out that the Bangladeshi outbreak was caused by a clone related to the South American lineage of the pathogen. Now that we know the enemy, we can proceed to put in place an informed response plan. It’s challenging but at least we know the nature of the pathogen – a first step in any response plan to a disease outbreak.
Which emerging diseases do you foresee having a large impact on food security in the future?
Obviously, any disease outbreak in the major food crops would be of immediate concern, but we shouldn’t neglect the smaller crops, which are so critical to agriculture in the developing world. This is one of the challenges of plant pathology: how to handle the numerous plants and their many pathogens.

European corn boreer. Image from Cornell University. Used under license CC BY 2.0.
As far as new problems, I view insect pests as being a particular challenge. Our basic understanding of insect–plant interactions is not as well developed as it is for microbial pathogens, and research has somewhat neglected the impact of plant immunity. The range of many insect pests is expanding because of climate change, and we are moving to ban many of the widely used insecticides. This is an area of research I would recommend for an early career scientist.
What advice would you give to a young researcher in this area?
Ask the right questions and look beyond the current trends. Think big. Be ambitious. Don’t shy away from embracing the latest technologies and methods. It’s important to work on real world systems. Thanks to technological advances, genomics, genome editing etc., the advantages of working on model systems are not as obvious as they were in the past.
How can we mitigate the risks to crops from plant diseases in the future?
My general take is to be suspicious of silver bullets. I like to say “Don’t bet against the pathogen”. I believe that for truly sustainable solutions, we need to continuously alter the control methods, for example by regularly releasing new resistant crop varieties. Only then we can keep up with rapidly evolving pathogens. One analogy would be the flu jab, which has a different formulation every year depending on the make-up of the flu virus population.

Potato with blight, caused by the oomycete Phytophthora infestans. Image credit: USDA. Used under license CC BY-ND 2.0.
Is there anything else you’d like to add?
I read that public and private funding of plant science is less than one tenth of biomedical research. Not a great state of affairs when one considers that we will add another two billion people to the planet in the next 30 years. As one of my colleagues once said: “medicine might save you one day; but plants keep you alive everyday”.
Following on from last week’s post, Now That’s What I Call Plant Science 2015, we bring you a year in Plant Science!

Image credit: Jean Weber. Used under license CC BY 2.0.
The year began with a surprising paper that turned our understanding of the phytohormone auxin on its head. Researchers in China and the USA created Arabidopsis knockout mutants of AUXIN BINDING PROTEIN 1 (ABP1), expecting them to fail to respond to auxin and have developmental defects, as previously seen in the abp1-1 knockdown mutant. Instead, these plants were indistinguishable from wild type plants, leading the authors to conclude that ABP1 is not required for auxin signaling or Arabidopsis development as previously believed.
Read the paper in PNAS: Auxin binding protein 1 (ABP1) is not required for either auxin signaling or Arabidopsis development.
A paper later in the year from the same authors found that the embryonic lethality of the abp1-1 mutant is actually caused by the off-target linked deletion of the adjacent BSM gene.
Read this paper in Nature Plants: Embryonic lethality of Arabidopsis abp1-1 is caused by deletion of the adjacent BSM gene.
The tale of ABP1 was examined in more detail on the GARNet blog, Weeding the Gems, which concluded: “In many ways this story is an excellent example of how science should work, where claims are independently tested to ensure that earlier experiments have been conducted or interpreted correctly.” Click here to read more.
A clever experiment from Germany led to a significant breakthrough in crop protection from insect pests.
When double-stranded RNA (dsRNA) is present within a eukaryotic cell, it is cleaved by the Dicer enzyme to form short interfering RNAs. These can bind to complementary RNA within a cell to target it for destruction, thus silencing the corresponding gene expression. This process is known as RNA interference (RNAi).
RNAi has previously been used to tackle insect herbivory by expressing insect-specific dsRNA in plants; however the protection has previously been incomplete. In this new study, published in Science, researchers produced dsRNA within chloroplasts, which do not have RNAi machinery. When dsRNA is expressed in the cytoplasm, the plant’s own Dicer enzyme breaks most of it down. When expressed in the chloroplasts, the dsRNA remained intact when eaten by insects, which proved much more effective at killing these pests.
Read the paper here: Full crop protection from an insect pest by expression of long double-stranded RNAs in plastids.
Another crop protection study followed in March, when researchers in China cloned the genetic locus in rice that confers broad-spectrum resistance to planthoppers – insect pests that cause the loss of billions of dollars of crops per year. Three lectin receptor kinase genes were found in rice cultivars from the Philippines, which enable plants to survive an infestation of insects. When cloned into a susceptible rice cultivar, these genes conferred resistance to two different planthopper species.
Understanding the genetic basis of resistance is very important as marker-assisted breeding and selection could be used to develop resistant rice varieties, and potentially utilized in other species of cereal.
Read the paper in Nature Biotechnology: A gene cluster encoding lectin receptor kinases confers broad-spectrum and durable insect resistance in rice
A European collaboration led to the development of 3DCellAtlas, a computational approach that semi-automatically identifies cell types in a developing 3D organ without the need for transgenic lineage markers. This program will enable the interpretation of dynamic organ growth and the spatial and temporal context of developmental cell divisions that produce the resultant plant. It could be integrated with growth in different conditions or with developmental mutants to examine exactly how these processes affect growth in 3D.
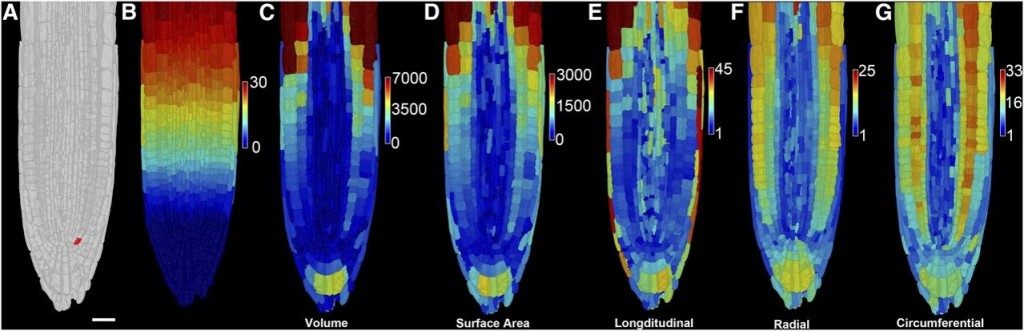
Image credit: Montenegro-Johnson et al., 2015. Digital Single-Cell Analysis of Plant Organ Development Using 3DCellAtlas. The Plant Cell, vol. 27 no. 4, 1018–1033.
A special issue of the Plant Biotechnology Journal was published in May, focusing on the amazing advances in molecular farming. While the entire issue is worth delving into, we were particularly intrigued by the review on moss-made pharmaceuticals, which outlines the rapid progress made in the field.
The model moss Physcomitrella patens has rapidly become one of the organisms of choice in biotechnology, with a fully sequenced genome and an outstanding toolbox for genome-engineering. The authors describe how moss-made pharmaceuticals can easily be produced while remaining remarkably more stable from batch to batch than cultured animal cells. The system is easily scalable, making their production highly cost effective, and safe. The first moss-made pharmaceuticals are currently in clinical trials, so keep an eye out for much more from this field over the next few years.
Read the review: Moss-made pharmaceuticals: from bench to bedside.
In June, US researchers discovered a new role for chloroplast stromules, protrusions that extend from the surface of all plastid types. The function of stromules has been difficult to determine, but this research, published in Developmental Cell, suggests that they may provide a mechanism by which plastid signals are conveyed to the nucleus. The paper shows that chloroplast stromules are induced by defense responses such as programmed cell death signaling, and that the stromules extend to form dynamic connections with the nucleus. The stromules may therefore aid in the amplification and/or transport of immune response signals into the nucleus.
Read the paper: Chloroplast Stromules Function during Innate Immunity.
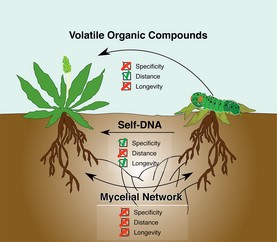
Image credit: Veresoglou et al., 2015. Self-DNA: a blessing in disguise? New Phytologist, vol. 207, no. 3, 488–490.
In late 2014 and early 2015, Italian researchers published a set of articles showing that extracellular self-DNA, DNA from conspecifics, could inhibit the growth of organisms from a wide range of taxa, including plants, bacteria, fungi and animals. Conversely, these organisms were not affected by extracellular DNA from other unrelated species.
In July, New Phytologist published a letter offering an interpretation of the data as it relates to plants. Plants could interpret extracellular self-DNA as an indicator of intraspecific competition (which seeds could use as a cue to remain dormant) or of a hostile environment that has already caused the death of conspecifics, signaling them to ramp up their pre-emptive immune response to increase survival after neighbors have been damaged or killed. There are still a lot of mechanisms and ecological effects to be investigated in this new field, but this letter suggests several interesting avenues to investigate.
Read the article: Self-DNA: a blessing in disguise?
Original research papers in New Phytologist:
Inhibitory effects of extracellular self-DNA: a general biological process?
A US study in August revealed a surprising degree of conservation in gene expression patterns across a wide range of plant taxa during root development. This was particularly interesting because the spikemoss Selaginella was shown to use many of the same genes as the evolutionarily distant angiosperms, despite the fossil record suggesting that roots evolved independently in these two lineages. Perhaps roots in these two groups evolved by independently recruiting the same developmental program, or perhaps by elaborating on a previously unknown proto-root that existed in the common ancestor of vascular plants.
Read the paper in The Plant Cell: Conserved Gene Expression Programs in Developing Roots from Diverse Plants.
Salt stress can significantly reduce the growth and yield of plants. Researchers in Germany identified two components of the cellulose synthase complex that directly interact with the microtubules and promote their dynamics, which interestingly were highly produced during salt stress conditions. During salt stress, cellulose microtubules depolymerize, however the newly discovered compounds, known as Companions of Cellulose Synthase, promote the reassembly of the microtubule to allow cellulose synthesis to continue.
Read the paper in Cell: A Mechanism for Sustained Cellulose Synthesis during Salt Stress
Throughout the year the GM debate in Europe reached several important milestones. In January the European Union (EU) changed its rules, giving individual countries more flexibility to decide for themselves whether or not to plant GM crops. In February, the UK Science and Technology Committee report stated that EU regulations preventing GM crops are not fit for purpose, and that they should be replaced with a trait-based system.
In October, EU member states revealed their stances on GM crops, with over half of Europe opting out of growing GM crops. Germany was the largest country to opt out of growing GM. The full list can be viewed here: Restrictions of geographical scope of GMO.
Read the news articles here:
EU changes rules on GM crop cultivation – January 2015
EU regulation on GM Organisms not ‘fit for purpose’ – February 2015
Half of Europe opts out of new GM crop scheme – October 2015

Image credit: University of Warwick, UK.
A collaboration between South African and UK scientists revealed how plants can use their circadian clock to pre-emptively boost their immune resistance at dawn, when fungal infection is most likely. Plants tend to decrease in susceptibility at dawn, but those with dysfunctional circadian clocks remained highly susceptible throughout the day. The research also showed that jasmonate signaling plays a crucial role in the circadian timing of resistance.
Read the article in The Plant Journal: Jasmonate signalling drives time-of-day differences in susceptibility of Arabidopsis to the fungal pathogen Botrytis cinerea.

Image credit: Guo & Liu., 2015. A single-nucleotide exon found in Arabidopsis. Scientific Reports, 5:18087.
Researchers in China published the surprising finding that a single-nucleotide exon exists in the APC11 gene in Arabidopsis. This is the smallest exon ever to be discovered before. The team used an elegant set of APC11-GFP constructs to show that intron splicing around the single-nucleotide exon is effective in both Arabidopsis and rice. This finding has implications for future genome annotations, which might reveal many more single-nucleotide exons.
Read the paper in Scientific Reports: A single-nucleotide exon found in Arabidopsis.
What a wonderful year of science! What new knowledge will 2016 bring?