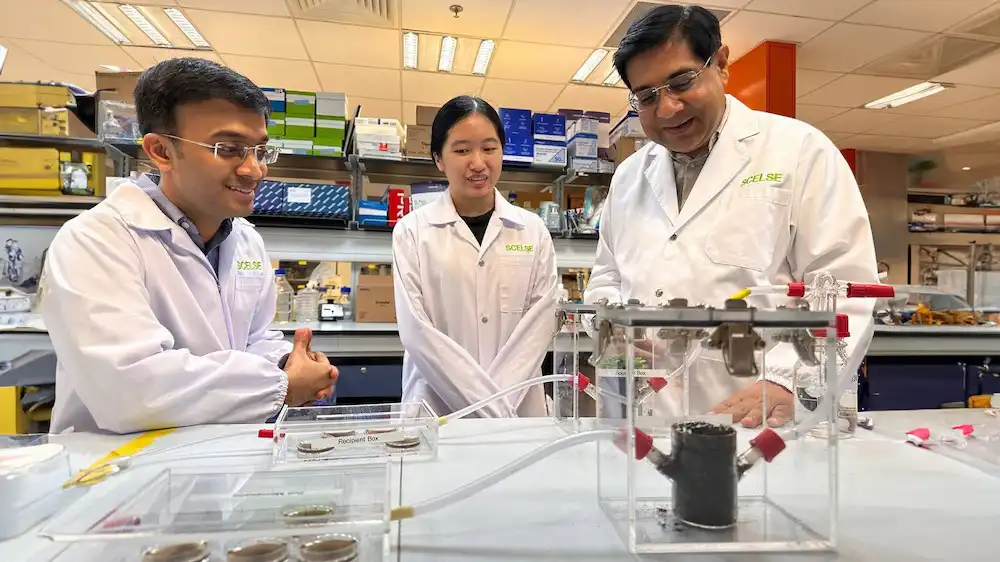
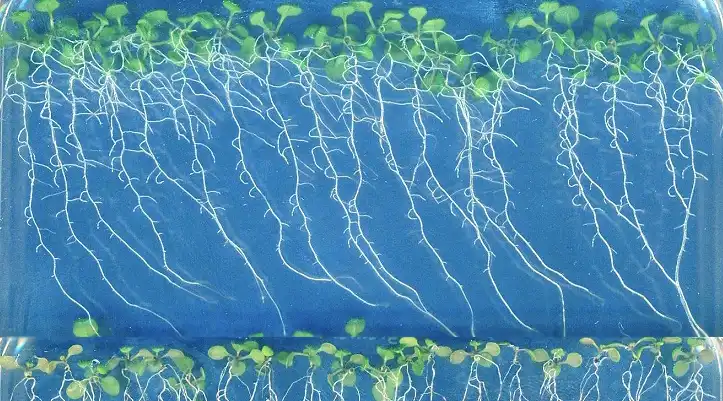
Plants emit odours for a variety of reasons, such as to communicate with each other, to deter herbivores or to respond to changing environmental conditions. An interdisciplinary team of researchers carried out a study to investigate how biodiversity influences the…
Read More