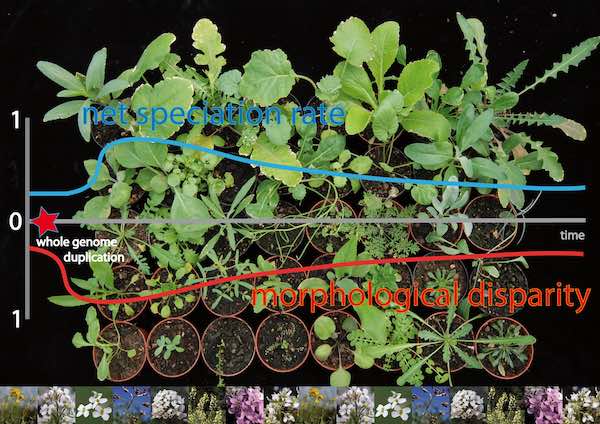
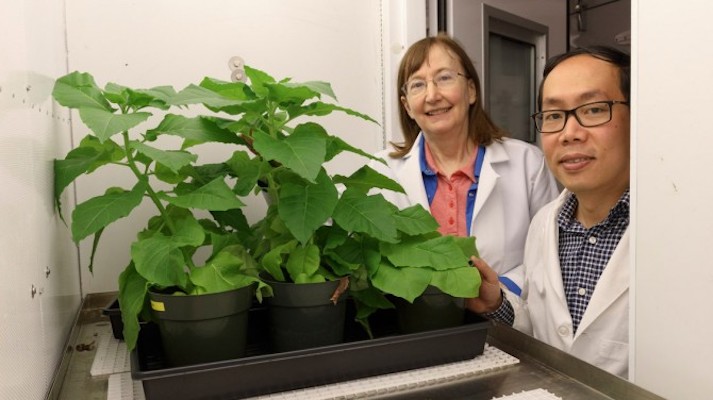
Genome duplications play a major role in the development of forms and structures of plant organisms and their changes across long periods of evolution. Biologists made this discovery in their research of the Brassicaceae family. To determine the scope of…
Read More