


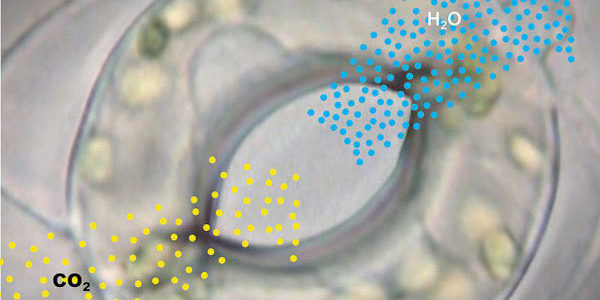
Plants face a dilemma in dry conditions: they have to seal themselves off to prevent losing too much water but this also limits their uptake of carbon dioxide. A sensory network assures that the plant strikes the right balance.
When water is scarce, plants can close their pores to prevent losing too much water. This allows them to survive even longer periods of drought, but with the majority of pores closed, carbon dioxide uptake is also limited, which impairs photosynthetic performance and thus plant growth and yield.
Plant accomplish a balancing act – navigating between drying out and starving in dry conditions – through an elaborate network of sensors. An international team of plant scientists led by Rainer Hedrich, a biophysicist from Julius-Maximilians-Universität (JMU) Würzburg in Bavaria, Germany, has now pinpointed these sensors. The results have been published in the journal Nature Plants.
When light is abundant, plants open the pores in their leaves to take in carbon dioxide (CO2) which they subsequently convert to carbohydrates in a process called photosynthesis. At the same time, a hundred times more water escapes through the microvalves than carbon dioxide flows in.
This is not a problem when there is enough water available, but when soils are parched in the middle of summer, the plant needs to switch to eco-mode to save water. Then plants will only open their pores to perform photosynthesis for as long as necessary to barely survive. Opening and closing the pores is accomplished through specialised guard cells that surround each pore in pairs. The units comprised of pores and guard cells are called stomata.
The guard cells must be able to measure the photosynthesis and the water supply to respond appropriately to changing environmental conditions. For this purpose, they have a receptor to measure the CO2 concentration inside the leaf. When the CO2 value rises sharply, this is a sign that the photosynthesis is not running ideally. Then the pores are closed to prevent unnecessary evaporation. Once the CO2 concentration has fallen again, the pores reopen.
The water supply is measured through a hormone. When water is scarce, plants produce abscisic acid (ABA), a key stress hormone, and set their CO2 control cycle to water saving mode. This is accomplished through guard cells which are fitted with ABA receptors. When the hormone concentration in the leaf increases, the pores close.
The JMU research team wanted to shed light on the components of the guard cell control cycles. For this purpose, they exposed Arabidopsis species to elevated levels of CO2 or ABA. They did so over several hours to trigger reactions at the level of the genes. Afterwards, the stomata were isolated from the leaves to analyse the respective gene expression profiles of the guard cells using bioinformatics techniques. For this task, the team took Tobias Müller and Marcus Dietrich on board, two bioinformatics experts at the University of Würzburg.
The two experts found out that the gene expression patterns differed significantly at high CO2 or ABA concentrations. Moreover, they noticed that excessive CO2 also caused the expression of some ABA genes to change. These findings led the researchers to take a closer look at the ABA signalling pathway. They were particularly interested in the ABA receptors of the PYR/PYL family (pyrabactin receptor and pyrabactin-like). Arabidopsis has 14 of these receptors, six of them in the guard cells.
“Why does a guard cell need as many as six receptors for a single hormone? To answer this question, we teamed up with Professor Pedro Luis Rodriguez from the University of Madrid, who is an expert in ABA receptors,” says Hedrich. Rodriguez’s team generated Arabidopsis mutants in which they could study the ABA receptors individually.
“This enabled us to assign each of the six ABA receptors a task in the network and identify the individual receptors which are responsible for the ABA- and CO2-induced closing of the stomata,” Peter Ache, a colleague of Hedrich‘s, explains.
“We conclude from the findings that the guard cells offset the current photosynthetic carbon fixation performance with the status of the water balance using ABA as the currency,” Hedrich explains. “When the water supply is good, our results indicate that the ABA receptors evaluate the basic hormonal balance as quasi ‘stress-free’ and keep the stomata open for CO2 supply. When water is scarce, the drought stress receptors recognise the elevated ABA level and make the guard cells close the stomata to prevent the plant from drying out.”
Next, the JMU researchers aim to study the special characteristics of the ABA and CO2 relevant receptors as well as their signalling pathways and components.
Read the paper: Nature Plants
Article source: UNIVERSITY OF WÜRZBURG
Image: Rainer Hedrich & Peter Ache / Universität Würzburg

Akiko Nakazaki, PhD student at Kyoto University
Kon-nichiwa! (Hello!) I am Akiko Nakazaki, a PhD student studying plant molecular cell biology at Kyoto University in Japan.
I’m interested in plant defense – specifically the glucosinolate-myrosinase defense system, which is specific to Brassicales such as Arabidopsis thaliana. Glucosinolates are a group of secondary metabolites stored in separate cells to myrosinases, the enzymes that break them down. Upon tissue damage, the glucosinolates and myrosinases are released from their cells and combine. The glucosinolates are hydrolyzed to volatile repellent compounds such as isothiocyanates and nitriles.
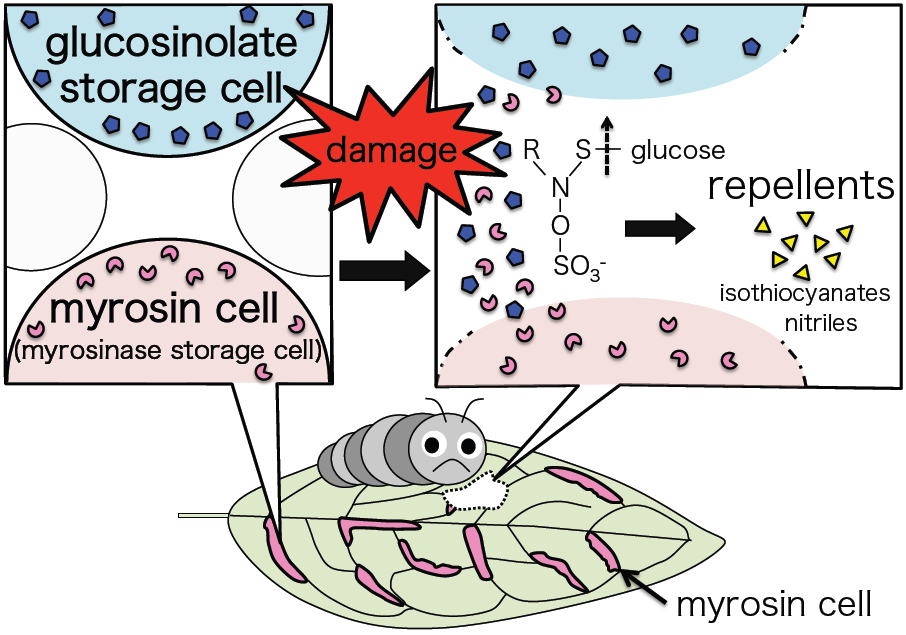
When damaged, cells containing glucosinolate and myrosinase are ruptured, releasing their contents. The glucosinolate is broken down by the myrosinase into volatile compounds that repel herbivores
I was impressed by this ingenious and rational survival strategy! I want to reveal this defense system at the cellular level, and am researching it in Arabidopsis thaliana by performing microscopic observations, bioassays with insects, and so on.
Are you interested in how PhD students from other countries spend their day in the laboratory? I am! Let me tell you about my typical day in the lab.
I wake up at 8:30am, and have morning coffee and toast for breakfast while reading a newspaper. Then, I get dressed and ride on my bicycle to the University. During the ride (about 10 minutes), I remind myself of the day’s schedule. I get to the lab at 10am and take my seat. All the members of the lab have their own desk and workbench. I turn on my computer and check my emails.
In the daylight, I basically do experiments and read papers. I start doing microscopic observations and lose track of time until I hear my stomach growling and realize that it is almost 2pm. I have lunch at the eating space in lab. In this room, there are always some lab members who are eating, discussing their research, playing social games, etc. After lunch, I report the result of my microscopic observations to my boss and we have a brief discussion about it.
Then, I return to my seat and realize the primers I ordered yesterday have arrived. I perform a PCR and prepare an agarose gel for electrophoresis. While I am waiting for the PCR to end, I search PubMed and Google Scholar for new papers to read. I load the PCR products to the gel and check that the PCR worked. In the evening, I allocate myself free time for doing more experiments, reading more papers, preparing research presentations, discussions, etc.
I’ve sought a more effective way to advance my research through trial and error. For example, when I started researching in the lab I was a little too ambitious, and planned my schedule too tightly. I sometimes felt tired and depressed when my research was not right on schedule, as is often the case. In these negative moods I couldn’t enjoy my work, so I adopted a schedule with more free time. Because of this change, I’ve come to be able to work flexibly and keep a positive frame of mind.
I’m home between 10pm and midnight. At home, I have a late dinner and take a good long soak in the bath (my favorite time of day!). I go to bed at 2am.
On weekends I enjoy playing badminton, learning traditional Japanese dance and shopping. I try to make plans without lab work as much as I can, however I’m not able to do avoid it sometimes when I am struggling to get new data before academic conferences and progress reports. Leaving the lab allows me to get rid of stress and feel refreshed for a healthy next week. Furthermore, I devise ways to work more efficiently on weekdays, because I am required to take time off at the weekends.
My boss always says, “It is important to value encounters with people and things.” It wasn’t until recently that I finally understood that message! I have found that experiments may not always work well, but when I look at it from a different angle, even experiments that haven’t gone the way I’d wanted could make me aware of something new and interesting. This awareness could also be brought about through discussions with others.
I am grateful for being able to receive this opportunity. Thank you.
Akiko Nakazaki is in the first year of her doctoral program in the Department of Botany, Graduate School of Science, Kyoto University, Japan.
Considering its weedy nature, Arabidopsis thaliana is a fussy little plant. This can be a pain – even tiny environmental fluctuations can have significant impacts on the physiology and development that many of us are investigating.
As silly as it sounds, my labmates and I have spent many months debating the best compost media to use when growing Arabidopsis for research. It began when our trusted compost supplier changed the formula of its peat-based compost, which stressed our plants and turned them a lovely shade of purple! The conversation has continued to develop as we learn about the different media used in other laboratories.
A new paper from Drake et al. at my university (University of Bristol, UK) has added a new depth to the debate, so I thought I’d bring it all to your attention and perhaps receive some other suggestions to consider!
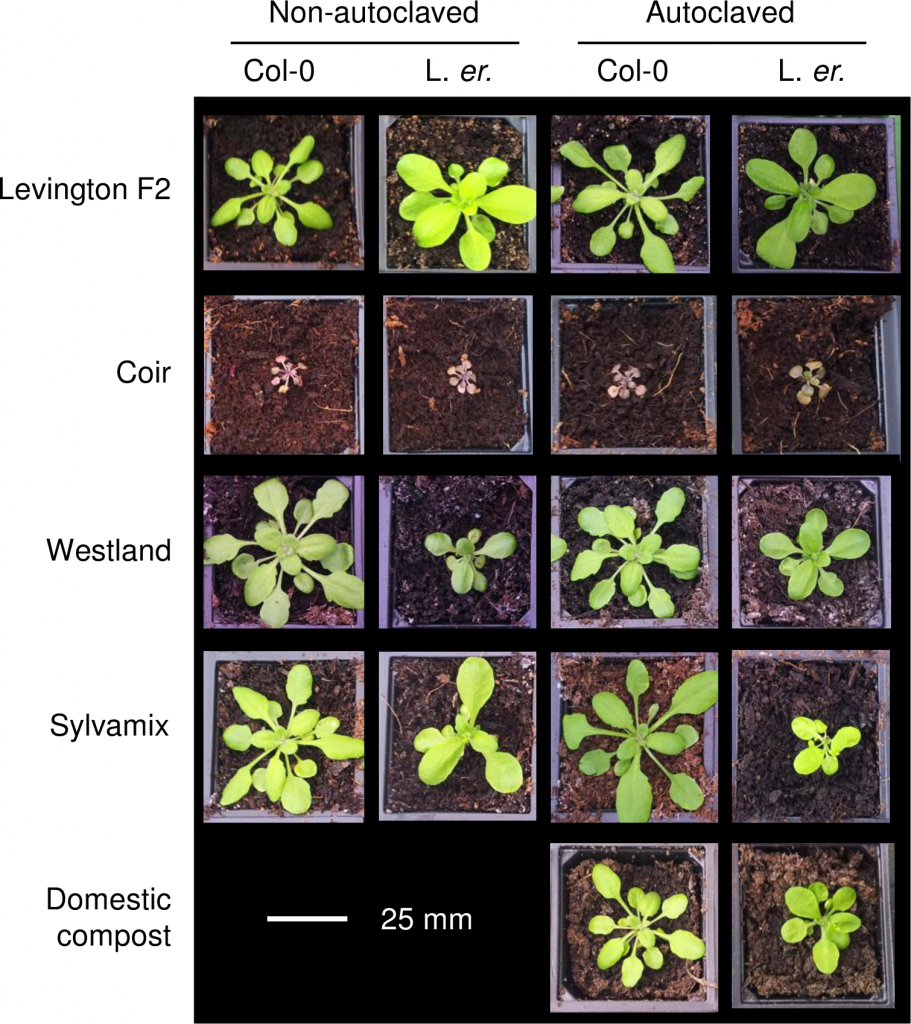

Arabidopsis growth on peat-based and peat-free growth media. Drake et al., 2016.
The experiment, led by Dr Antony Dodd, was designed to test whether peat-based composts could be replaced by alternatives in Arabidopsis research, in an attempt to reduce plant science’s use of unsustainable peat extraction. The researchers grew two ecotypes of Arabidopsis (Col-0 and Ler) on both autoclaved and non-autoclaved composts, including peat-based compost and some formed of coir, composted bark, wood-fiber, and a domestic compost.
In terms of reducing peat use, Arabidopsis unfortunately grew best on the peat-based growing media, although some vegetative traits were comparable in some peat-free composts.
This study caught my eye for another reason, however. We always sterilize our compost before growing Arabidopsis to reduce its contamination by fungi and insect pests; however, after learning that manganese toxicity can become a problem, we no longer autoclave it. As you can see in Boyd’s 1971 paper, manganese is converted to a more bioavailable form during the autoclave process, which can be toxic to plants.
Interestingly, Drake et al.’s research revealed no differences in Arabidopsis growth on autoclaved vs. non-autoclaved media, but I expect that in other environmental conditions the elevated manganese availability could become a problem. They did find that the autoclaved soil actually had more issues with mildew and algae, possibly because the natural microbiota had been killed and the compost was therefore easier to colonize.
One of the biggest issues our lab has with non-autoclaved soil is the presence of small insects, which can predate our precious plants. A potential alternative to autoclaving is to treat the media with insecticide, such as imidacloprid, a neonicotinoid. However, many labs have stopped using these pesticides; in 2010, Ford et al. showed that several neonicotinoids, including imidacloprid, induce salicylate-associated plant defense responses associated with enhanced stress tolerance, while in 2012, Cheng et al. found 225 genes were differentially expressed in rice plants treated with imidacloprid. In experiments designed to measure precise physiological responses, I’m not convinced that it’s a good idea to use these pesticides!
To avoid using autoclaves and insecticides, you could consider baking compost overnight at 60°C (140°F) to try and kill fungal spores and insects, freezing the media, and/or using biocontrols to tackle insect pests, such as nematodes or mites.
In the peat vs. non-peat debate, it looks as though peat-based media are still the frontrunners in terms of compost, but hydroponic systems are becoming more popular as a way of tightly controlling nutrient regimes and manipulating whole plants more easily. Check out this video from Associate Professor Matthew Gilliham (University of Adelaide, Australia) to learn more about the technique:
Following on from last week’s post, Now That’s What I Call Plant Science 2015, we bring you a year in Plant Science!

Image credit: Jean Weber. Used under license CC BY 2.0.
The year began with a surprising paper that turned our understanding of the phytohormone auxin on its head. Researchers in China and the USA created Arabidopsis knockout mutants of AUXIN BINDING PROTEIN 1 (ABP1), expecting them to fail to respond to auxin and have developmental defects, as previously seen in the abp1-1 knockdown mutant. Instead, these plants were indistinguishable from wild type plants, leading the authors to conclude that ABP1 is not required for auxin signaling or Arabidopsis development as previously believed.
Read the paper in PNAS: Auxin binding protein 1 (ABP1) is not required for either auxin signaling or Arabidopsis development.
A paper later in the year from the same authors found that the embryonic lethality of the abp1-1 mutant is actually caused by the off-target linked deletion of the adjacent BSM gene.
Read this paper in Nature Plants: Embryonic lethality of Arabidopsis abp1-1 is caused by deletion of the adjacent BSM gene.
The tale of ABP1 was examined in more detail on the GARNet blog, Weeding the Gems, which concluded: “In many ways this story is an excellent example of how science should work, where claims are independently tested to ensure that earlier experiments have been conducted or interpreted correctly.” Click here to read more.
A clever experiment from Germany led to a significant breakthrough in crop protection from insect pests.
When double-stranded RNA (dsRNA) is present within a eukaryotic cell, it is cleaved by the Dicer enzyme to form short interfering RNAs. These can bind to complementary RNA within a cell to target it for destruction, thus silencing the corresponding gene expression. This process is known as RNA interference (RNAi).
RNAi has previously been used to tackle insect herbivory by expressing insect-specific dsRNA in plants; however the protection has previously been incomplete. In this new study, published in Science, researchers produced dsRNA within chloroplasts, which do not have RNAi machinery. When dsRNA is expressed in the cytoplasm, the plant’s own Dicer enzyme breaks most of it down. When expressed in the chloroplasts, the dsRNA remained intact when eaten by insects, which proved much more effective at killing these pests.
Read the paper here: Full crop protection from an insect pest by expression of long double-stranded RNAs in plastids.
Another crop protection study followed in March, when researchers in China cloned the genetic locus in rice that confers broad-spectrum resistance to planthoppers – insect pests that cause the loss of billions of dollars of crops per year. Three lectin receptor kinase genes were found in rice cultivars from the Philippines, which enable plants to survive an infestation of insects. When cloned into a susceptible rice cultivar, these genes conferred resistance to two different planthopper species.
Understanding the genetic basis of resistance is very important as marker-assisted breeding and selection could be used to develop resistant rice varieties, and potentially utilized in other species of cereal.
Read the paper in Nature Biotechnology: A gene cluster encoding lectin receptor kinases confers broad-spectrum and durable insect resistance in rice
A European collaboration led to the development of 3DCellAtlas, a computational approach that semi-automatically identifies cell types in a developing 3D organ without the need for transgenic lineage markers. This program will enable the interpretation of dynamic organ growth and the spatial and temporal context of developmental cell divisions that produce the resultant plant. It could be integrated with growth in different conditions or with developmental mutants to examine exactly how these processes affect growth in 3D.
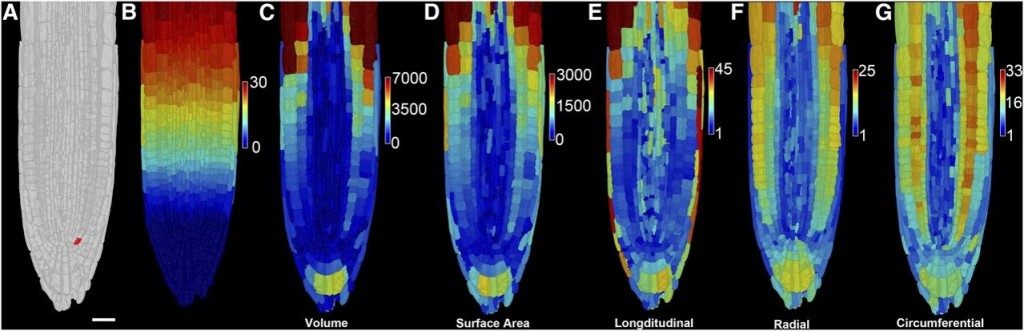
Image credit: Montenegro-Johnson et al., 2015. Digital Single-Cell Analysis of Plant Organ Development Using 3DCellAtlas. The Plant Cell, vol. 27 no. 4, 1018–1033.
A special issue of the Plant Biotechnology Journal was published in May, focusing on the amazing advances in molecular farming. While the entire issue is worth delving into, we were particularly intrigued by the review on moss-made pharmaceuticals, which outlines the rapid progress made in the field.
The model moss Physcomitrella patens has rapidly become one of the organisms of choice in biotechnology, with a fully sequenced genome and an outstanding toolbox for genome-engineering. The authors describe how moss-made pharmaceuticals can easily be produced while remaining remarkably more stable from batch to batch than cultured animal cells. The system is easily scalable, making their production highly cost effective, and safe. The first moss-made pharmaceuticals are currently in clinical trials, so keep an eye out for much more from this field over the next few years.
Read the review: Moss-made pharmaceuticals: from bench to bedside.
In June, US researchers discovered a new role for chloroplast stromules, protrusions that extend from the surface of all plastid types. The function of stromules has been difficult to determine, but this research, published in Developmental Cell, suggests that they may provide a mechanism by which plastid signals are conveyed to the nucleus. The paper shows that chloroplast stromules are induced by defense responses such as programmed cell death signaling, and that the stromules extend to form dynamic connections with the nucleus. The stromules may therefore aid in the amplification and/or transport of immune response signals into the nucleus.
Read the paper: Chloroplast Stromules Function during Innate Immunity.
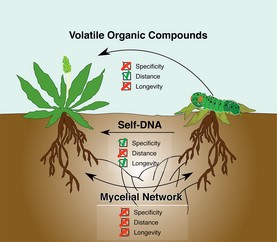
Image credit: Veresoglou et al., 2015. Self-DNA: a blessing in disguise? New Phytologist, vol. 207, no. 3, 488–490.
In late 2014 and early 2015, Italian researchers published a set of articles showing that extracellular self-DNA, DNA from conspecifics, could inhibit the growth of organisms from a wide range of taxa, including plants, bacteria, fungi and animals. Conversely, these organisms were not affected by extracellular DNA from other unrelated species.
In July, New Phytologist published a letter offering an interpretation of the data as it relates to plants. Plants could interpret extracellular self-DNA as an indicator of intraspecific competition (which seeds could use as a cue to remain dormant) or of a hostile environment that has already caused the death of conspecifics, signaling them to ramp up their pre-emptive immune response to increase survival after neighbors have been damaged or killed. There are still a lot of mechanisms and ecological effects to be investigated in this new field, but this letter suggests several interesting avenues to investigate.
Read the article: Self-DNA: a blessing in disguise?
Original research papers in New Phytologist:
Inhibitory effects of extracellular self-DNA: a general biological process?
A US study in August revealed a surprising degree of conservation in gene expression patterns across a wide range of plant taxa during root development. This was particularly interesting because the spikemoss Selaginella was shown to use many of the same genes as the evolutionarily distant angiosperms, despite the fossil record suggesting that roots evolved independently in these two lineages. Perhaps roots in these two groups evolved by independently recruiting the same developmental program, or perhaps by elaborating on a previously unknown proto-root that existed in the common ancestor of vascular plants.
Read the paper in The Plant Cell: Conserved Gene Expression Programs in Developing Roots from Diverse Plants.
Salt stress can significantly reduce the growth and yield of plants. Researchers in Germany identified two components of the cellulose synthase complex that directly interact with the microtubules and promote their dynamics, which interestingly were highly produced during salt stress conditions. During salt stress, cellulose microtubules depolymerize, however the newly discovered compounds, known as Companions of Cellulose Synthase, promote the reassembly of the microtubule to allow cellulose synthesis to continue.
Read the paper in Cell: A Mechanism for Sustained Cellulose Synthesis during Salt Stress
Throughout the year the GM debate in Europe reached several important milestones. In January the European Union (EU) changed its rules, giving individual countries more flexibility to decide for themselves whether or not to plant GM crops. In February, the UK Science and Technology Committee report stated that EU regulations preventing GM crops are not fit for purpose, and that they should be replaced with a trait-based system.
In October, EU member states revealed their stances on GM crops, with over half of Europe opting out of growing GM crops. Germany was the largest country to opt out of growing GM. The full list can be viewed here: Restrictions of geographical scope of GMO.
Read the news articles here:
EU changes rules on GM crop cultivation – January 2015
EU regulation on GM Organisms not ‘fit for purpose’ – February 2015
Half of Europe opts out of new GM crop scheme – October 2015

Image credit: University of Warwick, UK.
A collaboration between South African and UK scientists revealed how plants can use their circadian clock to pre-emptively boost their immune resistance at dawn, when fungal infection is most likely. Plants tend to decrease in susceptibility at dawn, but those with dysfunctional circadian clocks remained highly susceptible throughout the day. The research also showed that jasmonate signaling plays a crucial role in the circadian timing of resistance.
Read the article in The Plant Journal: Jasmonate signalling drives time-of-day differences in susceptibility of Arabidopsis to the fungal pathogen Botrytis cinerea.

Image credit: Guo & Liu., 2015. A single-nucleotide exon found in Arabidopsis. Scientific Reports, 5:18087.
Researchers in China published the surprising finding that a single-nucleotide exon exists in the APC11 gene in Arabidopsis. This is the smallest exon ever to be discovered before. The team used an elegant set of APC11-GFP constructs to show that intron splicing around the single-nucleotide exon is effective in both Arabidopsis and rice. This finding has implications for future genome annotations, which might reveal many more single-nucleotide exons.
Read the paper in Scientific Reports: A single-nucleotide exon found in Arabidopsis.
What a wonderful year of science! What new knowledge will 2016 bring?
With another year nearly over we recently put out a call for nominations for the Most Influential Plant Science Research of 2015. Suggestions flooded in, and we also trawled through our social media feeds to see which stories inspired the most discussion and engagement. It was fantastic to read about so much amazing research from around the world. Below are our top five, selected based on impact for the plant science research community, engagement on social media, and importance for both policy and potential end product/application.
Choosing the most inspiring stories was not an easy job. If you think we’ve missed something, please let us know in the comments below, or via Twitter! In the coming weeks we’ll be posting a 2015 Plant Science Round Up, which will include other exciting research that didn’t quite make the top five, so watch this space!

Sweet potato contains genes from bacteria making it a naturally occurring GM crop. Image from Mike Licht used under creative commons license 2.0
Scientists at the International Potato Center in Lima, Peru, found that 291 varieties of sweet potato actually contain bacterial genes. This technically means that sweet potato is a naturally occurring genetically modified crop! Alongside all the general discussion about GM regulations, particularly in parts of Europe where regulations about growing GM crops have been decentralized from Brussels to individual EU Member States, this story caused much discussion on social media when it was published in March of this year.
It is thought that ancestors of the modern sweet potato were genetically modified by bacteria in the soil some 8000 years ago. Scientists hypothesize that it was this modification that made consumption and domestication of the crop possible. Unlike the potato, sweet potato is not a tuber but a mere root. The bacteria genes are thought to be responsible for root swelling, giving it the fleshy appearance we recognize today.
This story is incredibly important, firstly because sweet potato is the world’s seventh most important food crop, so knowledge of its genetics and development are essential for future food supply. Secondly, Agrobacterium is frequently used by scientists to artificially genetically modify plants. Evidence that this process occurs in nature opens up the conversation about GM, the methods used in this technology, and the safety of these products for human consumption.
Read the original paper in PNAS here.
Our number two on the list also relates to genetic modification, this time focusing on methods. Regardless of whether or not we want to have genetically modified crops in our food supply, GM is a valuable tool used by researchers to advance knowledge of gene function at the genetic and phenotypic level. Therefore, systems of modification that make the process faster, cheaper, and more accurate provide fantastic opportunities for the plant science community to progress its understanding.
The Cas9 system is a method of genome editing that can make precise changes at specific locations in the genome relatively cheaply. This novel system uses small non-coding RNA to direct Cas9 nuclease to the DNA target site. This type of RNA is small and easy to program, providing a flexible and easily accessible system for genome editing.

Barley in the field. Image by Moldova_field used under creative commons license 2.0
Inheritance of genome modifications using Cas9 has previously been shown in the model plants, Arabidopsis and rice. However, the efficiency of this inheritance, and therefore potential application in crop plants has been questionable.
The breakthrough study published in November by researchers at The Sainsbury Laboratory and John Innes Centre both in Norwich, UK, demonstrated the mutation of two commercial crop plants, Barley and Brassica oleracea, using the Cas9 system and subsequent inheritance mutations.
This is an incredibly exciting development in the plant sciences and opens up many options in the future in terms of genome editing and plant science research.
Read the full paper in Genome Biology here.
Striga is a parasitic plant that mainly affects parts of Africa. It is a major threat to food crops such as rice and corn, leading to yield losses worth over 10 billion US dollars, and affecting over 100 million people.
Striga infects the host crop plant through its roots, depriving them of their nutrients and water. The plant hormone strigolactone, which is released by host plants, is known to induce Striga germination when host plants are nearby.
In a study published in August of this year the Striga receptors for this hormone, and the proteins responsible for striga germination were identified.
Striga plants are known to wither and die if they cannot find a host plant upon germination. Induction of early germination using synthetic hormones could therefore remove Striga populations before crops are planted. This work is vital in terms of regulating Striga populations in areas where they are hugely damaging to crop plants and people’s livelihoods.
Read the full study in Science here.

Striga, a parasitic plant. Also known as Witchweed. Image from the International Institute of Tropical Agriculture used under creative commons license 2.0
Resurrection plants are a unique group of flora that can survive extreme water shortages for months or even years. There are more than 130 varieties in the world, and many researchers believe that unlocking the genetic codes of drought-tolerant plants could help farmers working in increasingly hot and dry conditions.
During a drought, the plant acts like a seed, becoming so dry that it appears dead. But as soon as the rains come, the shriveled plant bursts ‘back to life’, turning green and robust in just a few hours.
In November, researchers from the Donald Danforth Plant Science Centre in Missouri, US, published the complete draft genome of Oropetium thomaeum, a resurrection grass species.
O. thomaeum is a small C4 grass species found in Africa and India. It is closely related to major food feed and bioenergy crops. Therefore this work represents a significant step in terms of understanding novel drought tolerance mechanisms that could be used in agriculture.
Read the full paper in Nature here.
Currently, one of the greatest challenges for ecologists is to quantify plant diversity and understand how this affects plant survival. For the last 500 years independent research groups around the world have collected this diversity data, which has made organization and collaboration difficult in the past.
Over the last 500 years, independent research groups have collected a wealth of diversity data. The Botanical Information and Ecology Network (BIEN) are collecting and collating these data together for the Americas using high performance computing (HPC) and data resources, via the iPlant Collaborative and the Texas Advanced Computing Center (TACC). This will allow researchers to draw on data right from the earliest plant collections up to the modern day to understand plant diversity.
There are approximately 120,000 plant species in North and South America, but mapping and determining the hotspots of species richness requires computationally intensive geographic range estimates. With supercomputing the BIEN group could generate and store geographic range estimates for plant species in the Americas.
It also gives ecologists the ability to document continental scale patterns of species diversity, which show where any species of plant might be found. These novel maps could prove a fantastic resource for ecologists working on diversity and conservation.
Read more about this story on the TACC website, here.