Researchers from the Okinawa Institute of Science and Technology Graduate University (OIST) have shed light on the reproductive role of ‘dark matter’ DNA – non-coding DNA sequences that previously seemed to have no function.
Their findings, published in Nature Communications, have revealed that a specific non-coding genomic region is essential for the proper development of the male and female reproductive organs in rice.
“Rice is one of the major global crops and is the staple food in many countries, including Japan,” said Dr. Reina Komiya, senior author of the research paper and associate researcher from the OIST Science and Technology Group. “Further research into how these genomic regions affect plant reproduction could potentially lead to increased productivity and more stable yields of rice.”
Many previous developmental studies have focused on genes – the sections of DNA that provide instructions for making proteins. But in complex creatures like plants and animals, a large fraction of the genome – typically between 90-98% – doesn’t actually code for proteins.
The vast expanse of this ‘junk DNA’ has long puzzled biologists, with many dubbing it the ‘dark matter’ of the genome. But recent research suggests that many of these non-coding genomic regions may have a function after all, giving rise to non-coding RNA.
Scientists have now identified numerous types of non-coding RNA, ranging from small molecules only 20-30 nucleotide bases in length to long molecules of over 200 nucleotides. Although studies show that non-coding RNA plays a vital role in the regulation of gene expression – the process where a gene’s instructions are used to make RNA or protein – the precise function of each specific non-coding RNA remains poorly understood.
Dr. Komiya is particularly interested in reproduction-specific RNAs. “These are non-coding RNAs that are produced as the reproductive system forms. I wanted to uncover what role they play in the development of stamens and pistils, the male and female reproductive organs in plants.”
Making mutants
In the study, Dr. Komiya’s group focused on a reproduction-specific microRNA – a major class of small non-coding RNAs – called microRNA2118.
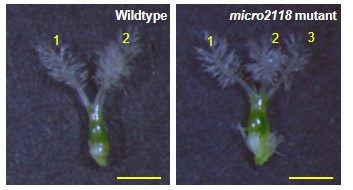
The scientists created mutant rice strains by deleting a region of the genome that contains multiple copies of the specific DNA sequence that gives rise to microRNA2118. They found that the mutant strains were sterile and showed abnormalities in the structure of the stamens and pistils.
The stamens in mutant strains contain anthers – the part of the stamen that houses pollen – that are shorter and more curled than in wildtype strains.
“This means that the role of microRNA2118 in the proper development of the stamens and pistils is essential for plant fertility,” said Dr. Komiya.
Revealing RNA and probing proteins
In order to delve deeper into how microRNA2118 controlled development of the anther, the scientists then identified which other molecules were affected by microRNA2118.
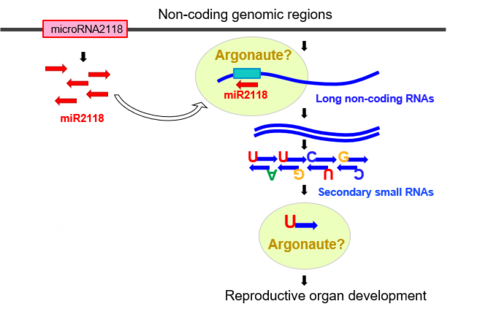
They found that microRNA2118 triggered the cleavage of long non-coding RNA, producing many tiny RNA molecules, called secondary small RNAs.
“Interestingly, these small RNAs were rich in uracil, one of the four nucleotide bases found in RNA, which is very unusual compared to other small RNAs,” said Dr. Komiya. “We hope to find out the exact function of these small RNAs – and whether this difference in nucleotide composition is important – in further research.”
The scientists also discovered that two Argonaute proteins that were only produced in the stamen were dependent on the presence of microRNA2118. Previous research has shown that Argonaute proteins team up with small RNAs to carry out many regulatory functions, such as silencing genes and cleaving RNA.
Dr. Komiya’s group therefore proposes that the Argonaute proteins may interact with microRNA2118 to trigger production of the secondary small RNAs. The proteins may also interact with the secondary small RNAs to silence specific regions of the genome. The team hopes to elucidate exactly how the Argonaute proteins and secondary small RNAs affect development of the plant reproductive system in further research.
“Reproduction is an important phenomenon of passing genetic information to the next generation and is essential for maintaining a stable yield supply. However, development of the reproductive system is complicated, and many aspects remain unknown,” concluded Dr. Komiya. “This study shows that non-coding RNAs, derived from regions of the genome that were thought to be non-functional, are vital for plant reproduction. Exploring non-coding RNAs further is an exciting and important area of research.”

Read the paper: Nature Communications
Article source: Okinawa Institute of Science and Technology (OIST) Graduate University
Author: Dani Ellenby
Images credit: Okinawa Institute of Science and Technology (OIST) Graduate University