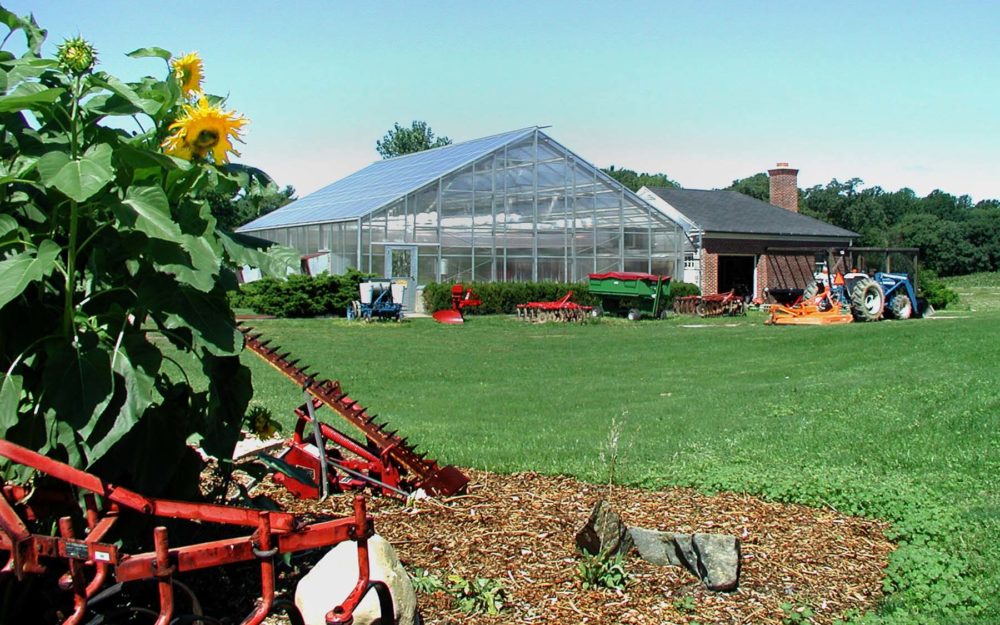
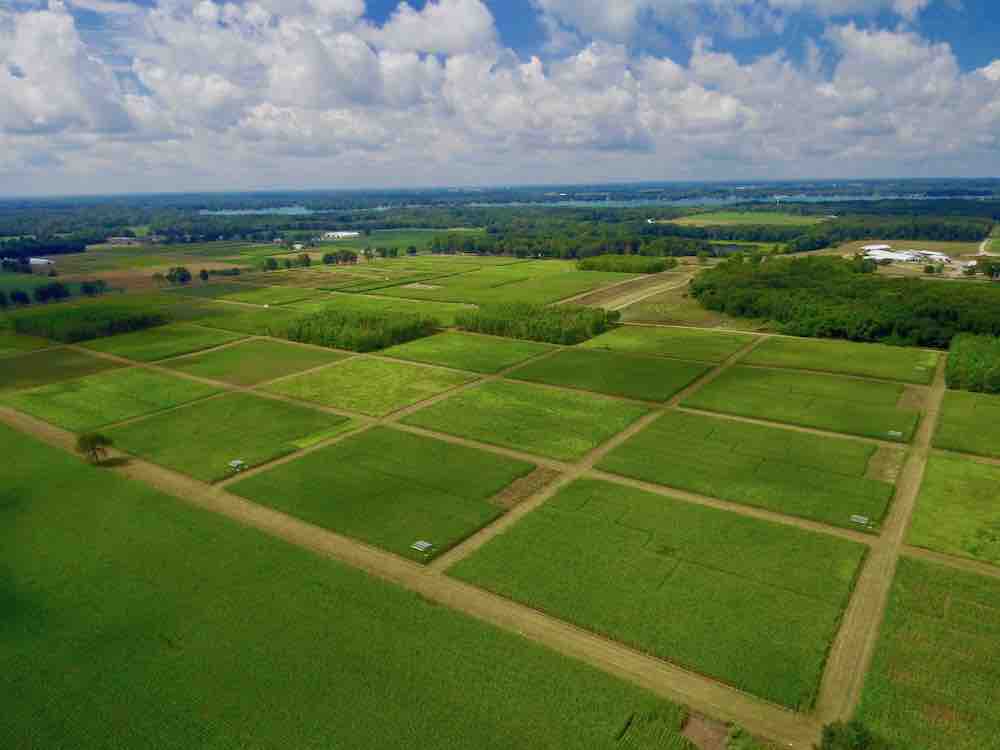




Grasslands make up more than 40% of the world’s ice-free land and have sustained humanity and thousands of other species for eons. In addition to providing food for cattle and sheep, grasslands are home to animals found nowhere else in the wild, such as the bison of North America’s prairies or the zebras and giraffes of the African savannas. Grasslands also can hold up to 30% of the world’s carbon, making them critical allies in the fight against climate change.
Climate change is causing grasslands to shift beneath our feet, putting these benefits at risk. Global change — which includes climate change, pollution and other widespread environmental alterations — is transforming grasslands and the plant species in them. A new study from researchers at Michigan State University shows what these changes to grassland plant communities look like, and reveal they are not always in ways scientists expect.
“Here in the Midwest, grasslands have been reduced to less than 1% of what they were at the time of European settlement and understanding what drove these changes is important to managing and restoring these systems” said Kay Gross, a plant ecologist at MSU’s Kellogg Biological Station, or KBS, and one of the authors of the study. “Our research at the KBS Long Term Ecological Research site and Allegan State Game Area had provided important information on these processes, but including our data into this larger synthesis reveals insights that are not apparent in site-specific research.”
The new paper, published in the Proceedings of the National Academy of Sciences, offers the most comprehensive evidence to date on how human activities are changing grassland plants.
The team looked at 105 grassland experiments around the world, including other sites from the National Science Foundation’s Long Term Ecological Research program and other research done at KBS. Each experiment tested at least one global change factor — such as rising carbon dioxide, hotter temperatures, extra nutrient pollution or drought. Some experiments looked at two or more of these factors. The team was led by Kimberly Komatsu, a grassland ecologist at the Smithsonian Environmental Research Center, and included researchers from around the world—including former KBS graduate students Emily Grman and Greg Houseman. Team members contributed data from a wide range of grasslands, and developed analyses to determine whether global change was altering the composition of grasslands, both in the total number and kinds of plant species present.
They discovered grasslands can be surprisingly tough — to a point. And it can take time for these changes to be detected. In general, grasslands resisted the effects of global change for the first decade of exposure. But after 10 years of exposure to a climate change factor, species began to shift. Half of the experiments lasting 10 years or more found a change in the total number of plant species, and nearly three-fourths found changes in the types of species. By contrast, only 20% of the experiments that lasted less than 10 years picked up any species changes at all. Experiments that examined three or more aspects of global change were also more likely to detect grassland transformation.
“I think grasslands are very, very resilient,” said Meghan Avolio, co-author and assistant professor of ecology at Johns Hopkins University. “But when conditions arrive that they do change, the change can be really important.”
To the scientists’ surprise, the identity of grassland species can change drastically, without altering the number of species. In half the plots where individual plant species changed, the total amount of species remained the same. In some plots, nearly all the species had changed.
For the team, this is a sign of hope that most grasslands could resist the experimentally induced global changes for at least 10 years. And that maybe grasslands are changing slowly enough that we can prevent catastrophic changes in the future.
However, time may not be on our side. In some experiments, the current pace of global change transformed even the “control plots” that were not exposed to experimentally higher global change pressures. Eventually, many of those plots looked the same as the experimental plots.
“Working collectively to understand how climate change is affecting grasslands is critical so that we can better restore and manage this important habitat that we and many other species depend on,” Gross said. “Long-term experiments and data sets are crucial for these efforts.”
Read the paper: Proceedings of the National Academy of Sciences
Article source: Michigan State University
Image credit: Kevin Kahmark, Michigan State University
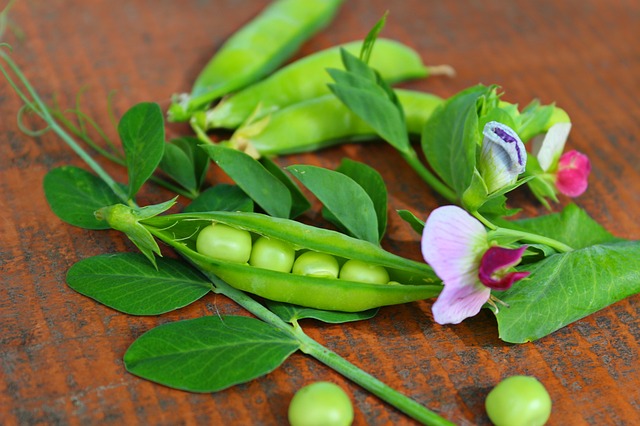
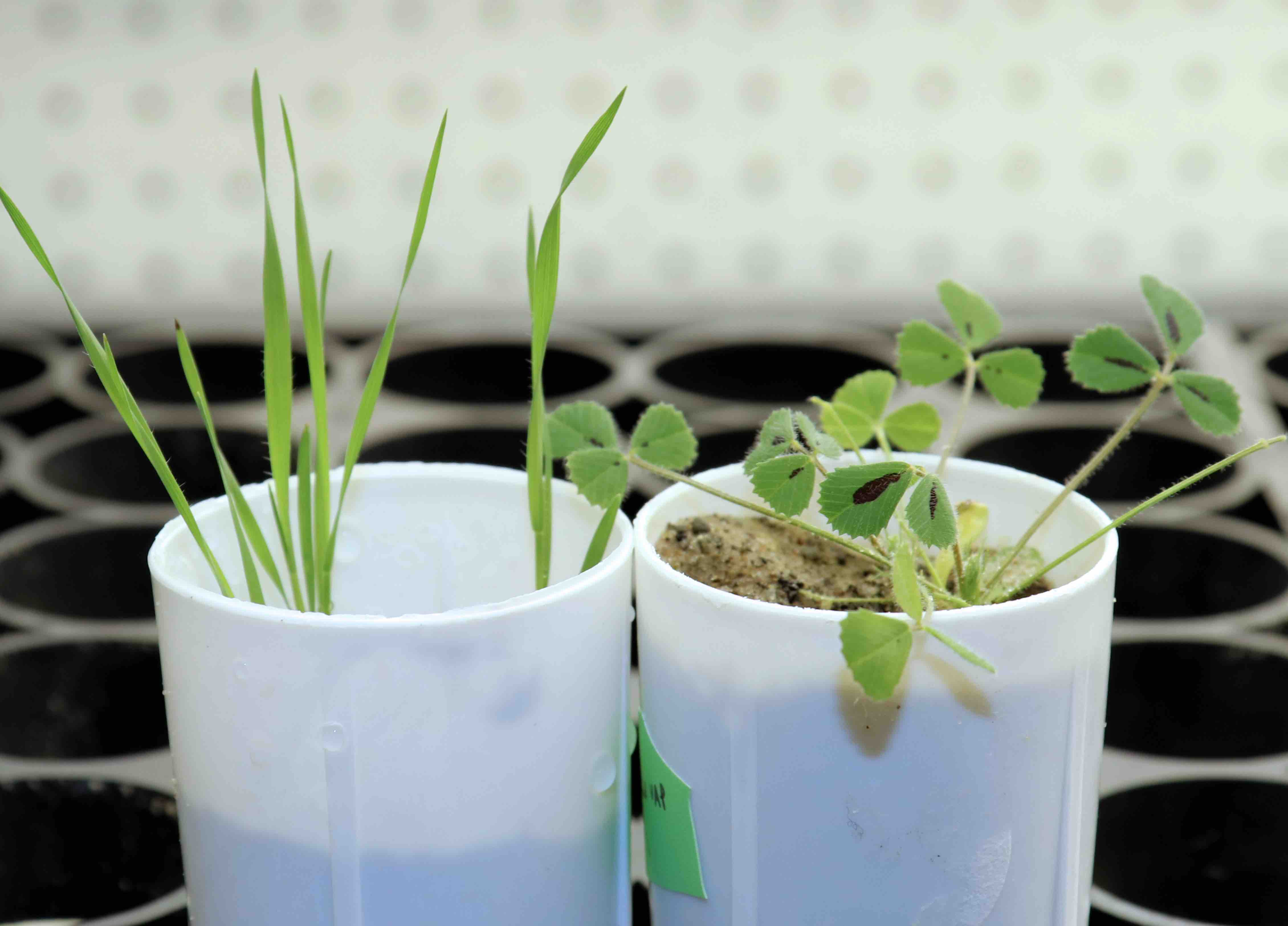




Over-fertilization of agricultural fields is a huge environmental problem. Excess phosphorus from fertilized cropland frequently finds its way into nearby rivers and lakes. A resulting boom of aquatic plant growth can cause oxygen levels in the water to plunge, leading to fish die-offs and other harmful effects.
Researchers from Boyce Thompson Institute have uncovered the function of a pair of plant genes that could help farmers improve phosphate capture, potentially reducing the environmental harm associated with fertilization.
The work was published in Nature Plants.
The discovery stems from Maria Harrison’s focus on plants’ symbiotic relationships with arbuscular mycorrhizal (AM) fungi. Harrison is the William H. Crocker Professor at BTI and an adjunct professor in Cornell University’s School of Integrative Plant Science.
AM fungi colonize plant roots, creating an interface where the plant trades fatty acids for phosphate and nitrogen. The fungi also can help plants recover from stressful conditions, such as periods of drought.
But feeding the AM fungi with fatty acids is costly, so plants don’t let this colonization go unchecked.
To discover how plants control the amount of fungal colonization, Harrison and Lena Müller, a postdoctoral scientist in her lab, looked at genes that encode short proteins called CLE peptides in the plants Medicago truncatula and Brachypodium distachyon.
CLE peptides are involved in cellular development and response to stress, and they are present throughout the plant kingdom, from green algae to flowering plants.
The researchers found that two of these CLE genes are key modulators of AM fungal symbiosis. One gene, called CLE53, reduces colonization rates once the roots have been colonized. Another gene, CLE33, reduces colonization rates when there is plenty of phosphate available to the plant.
“Being able to control fungal colonization levels in plant roots and maintain the symbiosis even in higher phosphate conditions might be useful to a farmer,” Harrison said. “For example, you may want the other beneficial effects of AM fungi, like nitrogen uptake and recovery from drought, as well as further uptake of phosphate”
“You might be able to achieve these benefits by altering the levels of these CLE peptides in the plants,” Harrison added.
Müller found that the CLE peptides act through a receptor protein called SUNN. In collaboration with Harro Bouwmeester and Kristyna Flokova of the University of Amsterdam, she found that the two CLE peptides modulate the plant’s synthesis of a compound called strigolactone.
Plant roots exude strigolactone into the soil, and the compound stimulates AM fungi to grow and colonize the root. Once the roots are colonized or there is plenty of phosphate, the CLE genes suppress the synthesis of strigolactone, thus reducing any further colonization by the fungi.
“In the early 2000’s, researchers found that plants had a way to measure and then reduce colonization,” Müller said. “But until now, nobody really understood the molecular mechanism of that dynamic.”
The researchers’ next steps will include figuring out the molecules that turn on the CLE genes in response to colonization and high phosphate levels.
Müller also plans to compare the two CLE peptides from this study with additional CLE peptides that have different functions.
“The CLE peptides are all so similar but they have completely different functions,” Müller said. “It will be very interesting to see why that is.”
Read the paper: Nature Plants
Article source: Boyce Thompson Institute
Image: Boyce Thompson Institute
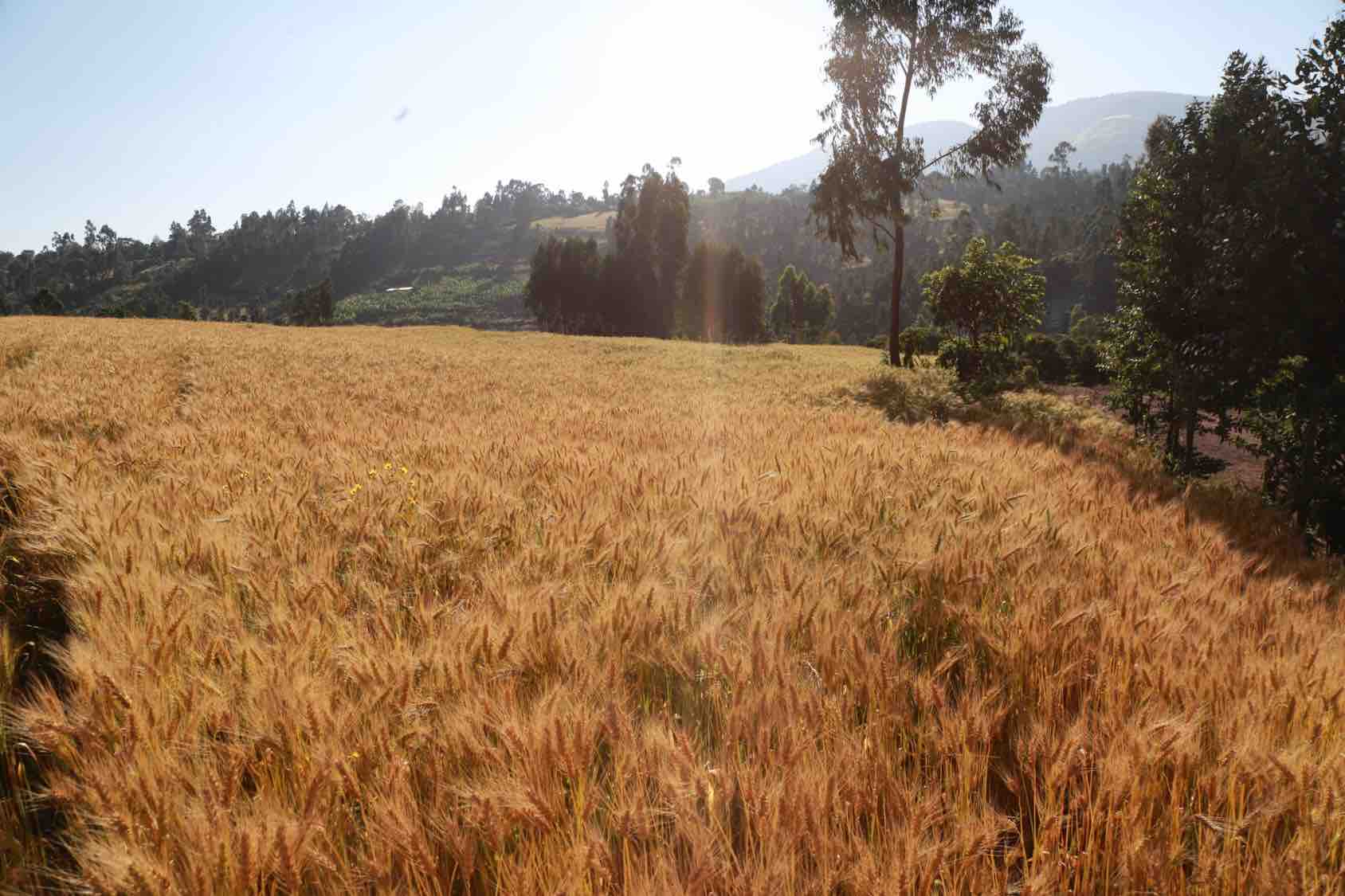

Scientists identified significant new chromosomal regions for wheat yield and disease resistance, which will speed up global breeding efforts.
Using the full wheat genome map published in 2018, combined with data from field testing of wheat breeding lines in multiple countries, an international team of scientists has identified significant new chromosomal regions for wheat yield and disease resistance and created a freely-available collection of genetic information and markers for more than 40,000 wheat lines.
Reported in Nature Genetics, the results will speed up global efforts to breed more productive and climate-resilient varieties of bread wheat, a critical crop for world food security that is under threat from rising temperatures, rapidly-evolving fungal pathogens, and more frequent droughts, according to Philomin Juliana, wheat scientist at the International Maize and Wheat Improvement Center (CIMMYT) and first author of the new study.
“This work directly connects the wheat genome reference map with wheat lines and extensive field data from CIMMYT’s global wheat breeding network,” said Juliana. “That network in turn links to over 200 breeding programs and research centers worldwide and contributes to yield and other key traits in varieties sown on nearly half the world’s wheat lands.”
The staple food for more than 2.5 billion people, wheat provides 20% of human dietary calories and protein worldwide and is critical for the nutrition and food security of hundreds of millions of poor persons in regions such as North Africa and South Asia.
“Farmers and societies today face new challenges to feed rising and rapidly-urbanizing populations, and wheat epitomizes the issues,” said Ravi Singh, CIMMYT wheat breeder and corresponding author of the study. “Higher temperatures are holding back yields in major wheat-growing areas, extreme weather events are common, crop diseases are spreading and becoming more virulent, and soil and water are being depleted.”
Juliana said the study results help pave the way to apply genomic selection, an approach that has transformed dairy cow husbandry, for more efficient wheat breeding.
“Molecular markers are getting cheaper to use; meanwhile, it’s very costly to do field testing and selection involving many thousands of wheat plants over successive generations,” Juliana said. “Genome-wide marker-based selection can help breeders to precisely identify good lines in early breeding generations and to test plantlets in greenhouses, thereby complementing and streamlining field testing.”
The new study found that genomic selection could be particularly effective in breeding for wheat end-use quality and for resistance to stem rust disease, whose causal pathogen has been evolving and spreading in the form of highly-virulent new races.
The new study also documents the effectiveness of the global public breeding efforts by CIMMYT and partners, showing that improved wheat varieties from this work have accumulated multiple gene variants that favor higher yields, according to Hans-Joachim Braun, director of CIMMYT’s global wheat program.
“This international collaboration, which is the world’s largest publicly-funded wheat breeding program, benefits farmers worldwide and offers high-quality wheat lines that are released directly to farmers in countries, such as Afghanistan, that are unable to run a full-fledged wheat breeding program,”
Braun explained.
The study results are expected to support future gene discovery, molecular breeding, and gene editing in wheat, Braun said.
Together with more resource-efficient cropping systems, high-yielding and climate-resilient wheat varieties will constitute a key component of the sustainable intensification of food production described in Strategy 3 of the recent EAT-Lancet Commission recommendations to transform the global food system. Large-scale genomics will play a key role in developing these varieties and staying ahead of climate- and disease-related threats to food security.
Read the paper: Nature Genetics
Article source: CIMMYT
Image: Apollo Habtamu/CIMMYT
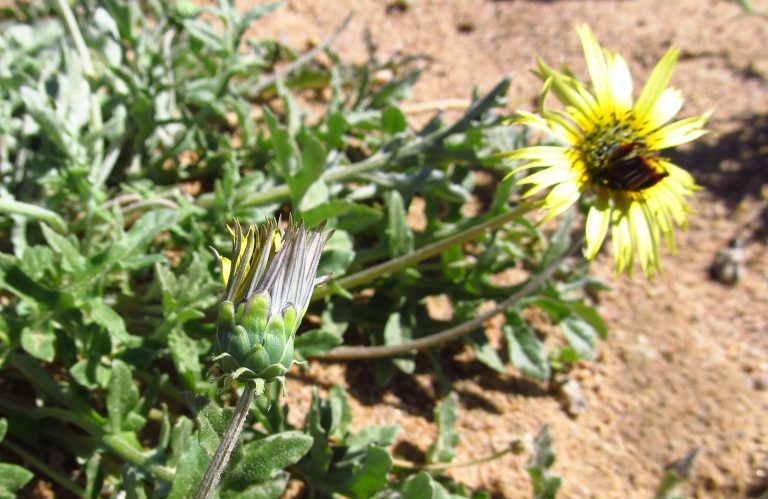

Species of daisy that close their flowers at night, produce colour in their exposed lower petals that makes them harder to spot for herbivores, reducing herbivory rates of flowers. The findings are presented in the British Ecological Society journal Functional Ecology.
Researchers from Stellenbosch University, South Africa found that tortoises, one of the main herbivores of the daisies, were unable to distinguish the lower petal surfaces against a green leaf background. Tortoises prefer to eat protein-rich flowers over leaves, but when confronted with closed flowers, they showed no preference between them.
When the researchers modelled the colours of the lower petal surfaces in the vision of other herbivores, they also found these colours to be indistinguishable from leaves.
In contrast, species of daisy that do not close at night produced the same colouration on their lower petals as the upper petals exposed to pollinators.
Plants face an evolutionary conflict between having flowers that attract pollinators while avoiding herbivores. Often plants defend themselves chemically, but this can have adverse effects on pollination.
“When plants defend their flowers chemically, the pollination interactions can be negatively influenced. Our study shows a novel way in which flowers can avoid herbivores, without compromising pollination interactions.
– says Dr. Jurene Kemp, lead author of the study.
“These flowers can potentially circumvent the conflict of attracting both pollinators and herbivores by producing attractive colours on the surfaces that are exposed to pollinators (when flowers are open) and cryptic colours that are exposed when herbivores are active (when flowers are closed).”
In Namaqualand, South Africa, where the research took place, daises bloom annually in a spring flowering. This makes preserving flowers, responsible for reproduction, particularly important.
The researchers examined the colouration of 77 Asteraceae species, modelling how they appear in the visual systems of chameleons, horses and goats as proxies for tortoises and larger herbivores in the area, like springbok. They then tested the preferences of real tortoises with both open and closed flowers against leaf backgrounds.
Not all Asteraceae species that close their flowers had cryptically coloured lower petal surfaces, but in the experiments, the tortoises did not readily eat these flowers. Dr. Kemp said, “One interesting question would be to test whether non-cryptic flowers have chemical defences, and whether these chemical defences are absent in the cryptic flowers.”
On further research Dr. Kemp said “Unfortunately, we could only do this using one plant family in one botanical region, it would be great to see if other plant species also use colour to avoid herbivores.”
The researchers would also have liked to use larger herbivores such as springboks in their behavioural experiments, but Dr. Kemp adds that “this was practically not possible.”
Read the paper: Functional Ecology
Article source: British Ecological society Press Office
Image: Jurene Kemp