


Another fantastic year of discovery is over – read on for our 2016 plant science top picks!

A Zostera marina meadow in the Archipelago Sea, southwest Finland. Image credit: Christoffer Boström (Olsen et al., 2016. Nature).
The year began with the publication of the fascinating eelgrass (Zostera marina) genome by an international team of researchers. This marine monocot descended from land-dwelling ancestors, but went through a dramatic adaptation to life in the ocean, in what the lead author Professor Jeanine Olsen described as, “arguably the most extreme adaptation a terrestrial… species can undergo”.
One of the most interesting revelations was that eelgrass cannot make stomatal pores because it has completely lost the genes responsible for regulating their development. It also ditched genes involved in perceiving UV light, which does not penetrate well through its deep water habitat.
Read the paper in Nature: The genome of the seagrass Zostera marina reveals angiosperm adaptation to the sea.
BLOG: You can find out more about the secrets of seagrass in our blog post.
Plants are known to form new organs throughout their lifecycle, but it was not previously clear how they organized their cell development to form the right shapes. In February, researchers in Germany used an exciting new type of high-resolution fluorescence microscope to observe every individual cell in a developing lateral root, following the complex arrangement of their cell division over time.
Using this new four-dimensional cell lineage map of lateral root development in combination with computer modelling, the team revealed that, while the contribution of each cell is not pre-determined, the cells self-organize to regulate the overall development of the root in a predictable manner.
Watch the mesmerizing cell division in lateral root development in the video below, which accompanied the paper:
Read the paper in Current Biology: Rules and self-organizing properties of post-embryonic plant organ cell division patterns.
In March, a Spanish team of researchers revealed how the anti-wilting molecular machinery involved in preserving cell turgor assembles in response to drought. They found that a family of small proteins, the CARs, act in clusters to guide proteins to the cell membrane, in what author Dr. Pedro Luis Rodriguez described as “a kind of landing strip, acting as molecular antennas that call out to other proteins as and when necessary to orchestrate the required cellular response”.
Read the paper in PNAS: Calcium-dependent oligomerization of CAR proteins at cell membrane modulates ABA signaling.
*If you’d like to read more about stress resilience in plants, check out the meeting report from the Stress Resilience Forum run by the GPC in coalition with the Society for Experimental Biology.*
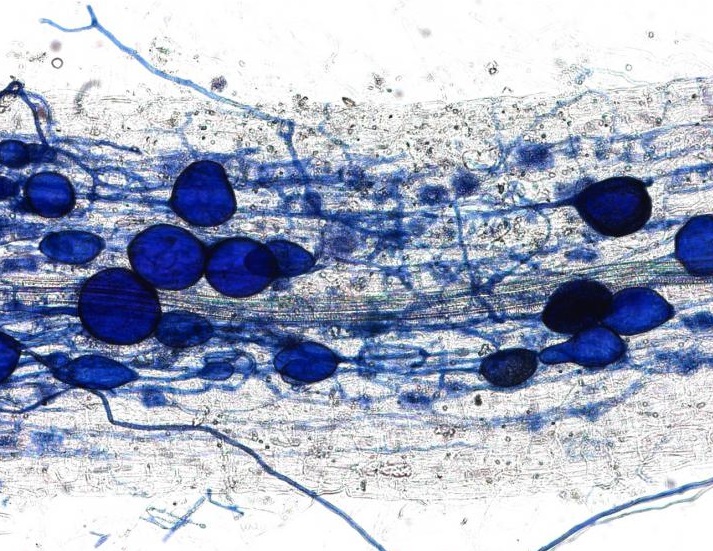
This plant root is infected with arbuscular mycorrhizal fungi. Image credit: University of Zurich.
In April, we received an amazing insight into the ‘decision-making ability’ of plants when a Swiss team discovered that plants can punish mutualist fungi that try to cheat them. In a clever experiment, the researchers provided a plant with two mutualistic partners; a ‘generous’ fungus that provides the plant with a lot of phosphates in return for carbohydrates, and a ‘meaner’ fungus that attempts to reduce the amount of phosphate it ‘pays’. They revealed that the plants can starve the meaner fungus, providing fewer carbohydrates until it pays its phosphate bill.
Author Professor Andres Wiemsken explains: “The plant exploits the competitive situation of the two fungi in a targeted manner, triggering what is essentially a market-based process determined by cost and performance”.
Read the paper in Ecology Letters: Options of partners improve carbon for phosphorus trade in the arbuscular mycorrhizal mutualism.
The transition of ancient plants from water onto land was one of the most important events in our planet’s evolution, but required a massive change in plant biology. Suddenly plants risked drying out, so had to develop new ways to survive drought.
In May, an international team discovered a key gene in moss (Physcomitrella patens) that allows it to tolerate dehydration. This gene, ANR, was an ancient adaptation of an algal gene that allowed the early plants to respond to the drought-signaling hormone ABA. Its evolution is still a mystery, though, as author Dr. Sean Stevenson explains: “What’s interesting is that aquatic algae can’t respond to ABA: the next challenge is to discover how this hormone signaling process arose.”
Read the paper in The Plant Cell: Genetic analysis of Physcomitrella patens identifies ABSCISIC ACID NON-RESPONSIVE, a regulator of ABA responses unique to basal land plants and required for desiccation tolerance.
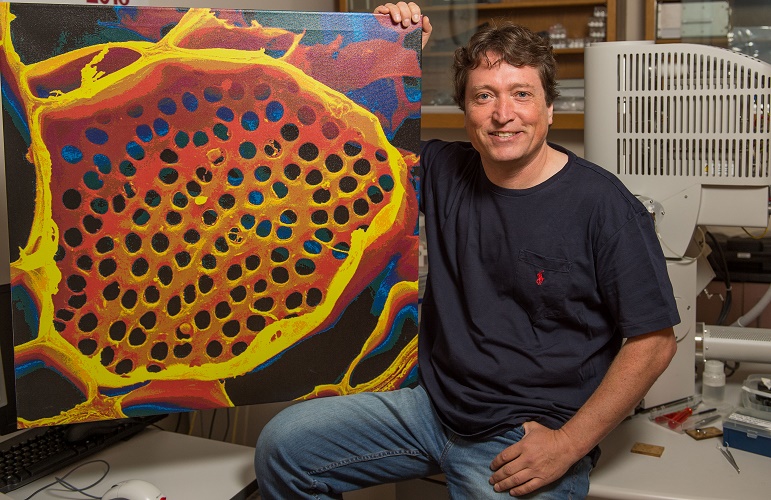
Professor Michael Knoblauch shows off a microscope image of phloem tubes. Image credit: Washington State University.
Sometimes revisiting old ideas can pay off, as a US team revealed in June. In 1930, Ernst Münch hypothesized that transport through the phloem sieve tubes in the plant vascular tissue is driven by pressure gradients, but no-one really knew how this would account for the massive pressure required to move nutrients through something as large as a tree.
Professor Michael Knoblauch and colleagues spent decades devising new methods to investigate pressures and flow within phloem without disrupting the system. He eventually developed a suite of techniques, including a picogauge with the help of his son, Jan, to measure tiny pressure differences in the plants. They found that plants can alter the shape of their phloem vessels to change the pressure within them, allowing them to transport sugars over varying distances, which provided strong support for Münch flow.
Read the paper in eLife: Testing the Münch hypothesis of long distance phloem transport in plants.
BLOG: We featured similar work (including an amazing video of the wound response in sieve tubes) by Knoblauch’s collaborator, Dr. Winfried Peters, on the blog – read it here!
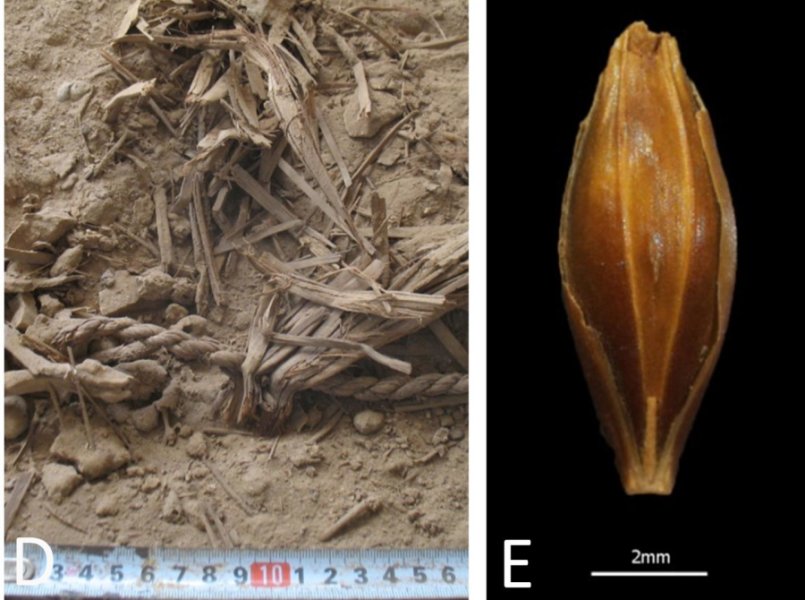

Preserved remains of rope, seeds, reeds and pellets (left), and a desiccated barley grain (right) found at Yoram Cave in the Judean Desert. Credit: Uri Davidovich and Ehud Weiss.
In July, an international and highly multidisciplinary team published the genome of 6,000-year-old barley grains excavated from a cave in Israel, the oldest plant genome reconstructed to date. The grains were visually and genetically very similar to modern barley, showing that this crop was domesticated very early on in our agricultural history. With more analysis ongoing, author Dr. Verena Schünemann predicts that “DNA-analysis of archaeological remains of prehistoric plants will provide us with novel insights into the origin, domestication and spread of crop plants”.
Read the paper in Nature Genetics: Genomic analysis of 6,000-year-old cultivated grain illuminates the domestication history of barley.
BLOG: We interviewed Dr. Nils Stein about this fascinating work on the blog – click here to read more!
Another exciting cereal paper was published in August, when an Australian team revealed that C4 photosynthesis occurs in wheat seeds. Like many important crops, wheat leaves perform C3 photosynthesis, which is a less efficient process, so many researchers are attempting to engineer the complex C4 photosynthesis pathway into C3 crops.
This discovery was completely unexpected, as throughout its evolution wheat has been a C3 plant. Author Professor Robert Henry suggested: “One theory is that as [atmospheric] carbon dioxide began to decline, [wheat’s] seeds evolved a C4 pathway to capture more sunlight to convert to energy.”
Read the paper in Scientific Reports: New evidence for grain specific C4 photosynthesis in wheat.
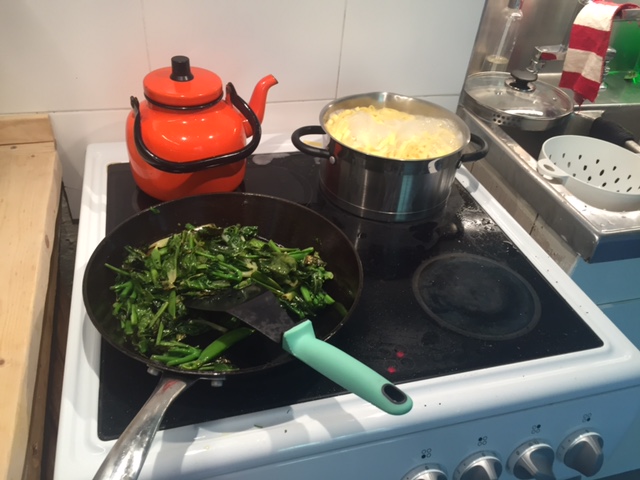

Professor Stefan Jansson cooks up “Tagliatelle with CRISPRy fried vegetables”. Image credit: Stefan Jansson.
September marked an historic event. Professor Stefan Jansson cooked up the world’s first CRISPR meal, tagliatelle with CRISPRy fried vegetables (genome-edited cabbage). Jansson has paved the way for CRISPR in Europe; while the EU is yet to make a decision about how CRISPR-edited plants will be regulated, Jansson successfully convinced the Swedish Board of Agriculture to rule that plants edited in a manner that could have been achieved by traditional breeding (i.e. the deletion or minor mutation of a gene, but not the insertion of a gene from another species) cannot be treated as a GMO.
Read more in the Umeå University press release: Umeå researcher served a world first (?) CRISPR meal.
BLOG: We interviewed Professor Stefan Jansson about his prominent role in the CRISPR/GM debate earlier in 2016 – check out his post here.
*You may also be interested in the upcoming meeting, ‘New Breeding Technologies in the Plant Sciences’, which will be held at the University of Gothenburg, Sweden, on 7-8 July 2017. The workshop has been organized by Professor Jansson, along with the GPC’s Executive Director Ruth Bastow and Professor Barry Pogson (Australian National University/GPC Chair). For more info, click here.*
Phytochromes help plants detect day length by sensing differences in red and far-red light, but a UK-Germany research collaboration revealed that these receptors switch roles at night to become thermometers, helping plants to respond to seasonal changes in temperature.
Dr Philip Wigge explains: “Just as mercury rises in a thermometer, the rate at which phytochromes revert to their inactive state during the night is a direct measure of temperature. The lower the temperature, the slower phytochromes revert to inactivity, so the molecules spend more time in their active, growth-suppressing state. This is why plants are slower to grow in winter”.
Read the paper in Science: Phytochromes function as thermosensors in Arabidopsis.
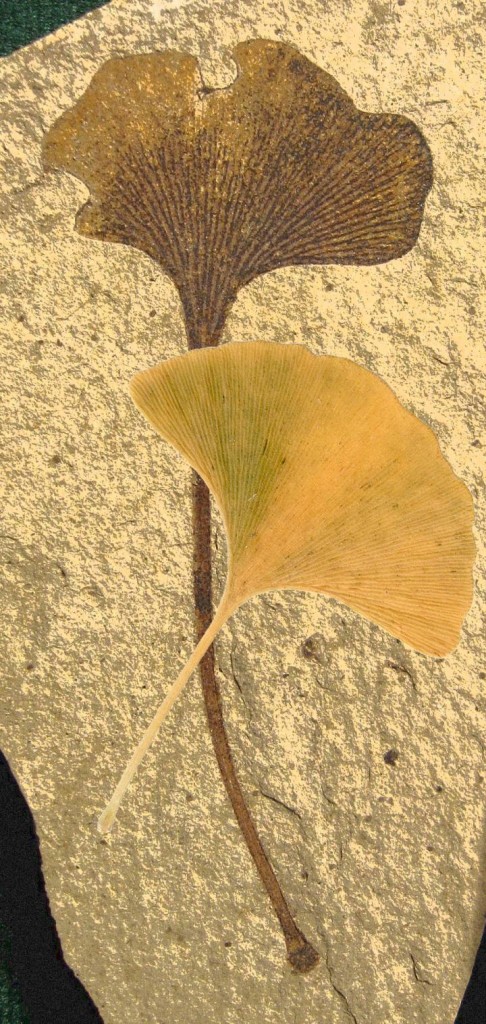

A fossil ginkgo (Ginkgo biloba) leaf with its modern counterpart. Image credit: Gigascience.
In November, a Chinese team published the genome of Ginkgo biloba¸ the oldest extant tree species. Its large (10.6 Gb) genome has previously impeded our understanding of this living fossil, but researchers will now be able to investigate its ~42,000 genes to understand its interesting characteristics, such as resistance to stress and dioecious reproduction, and how it remained almost unchanged in the 270 million years it has existed.
Author Professor Yunpeng Zhao said, “Such a genome fills a major phylogenetic gap of land plants, and provides key genetic resources to address evolutionary questions [such as the] phylogenetic relationships of gymnosperm lineages, [and the] evolution of genome and genes in land plants”.
Read the paper in GigaScience: Draft genome of the living fossil Ginkgo biloba.
The year ended with another fascinating discovery from a Danish team, who used fluorescent tags and microscopy to confirm the existence of metabolons, clusters of metabolic enzymes that have never been detected in cells before. These metabolons can assemble rapidly in response to a stimulus, working as a metabolic production line to efficiently produce the required compounds. Scientists have been looking for metabolons for 40 years, and this discovery could be crucial for improving our ability to harness the production power of plants.
Read the paper in Science: Characterization of a dynamic metabolon producing the defense compound dhurrin in sorghum.
Another amazing year of science! We’re looking forward to seeing what 2017 will bring!
P.S. Check out 2015 Plant Science Round Up to see last year’s top picks!
Following on from last week’s post, Now That’s What I Call Plant Science 2015, we bring you a year in Plant Science!

Image credit: Jean Weber. Used under license CC BY 2.0.
The year began with a surprising paper that turned our understanding of the phytohormone auxin on its head. Researchers in China and the USA created Arabidopsis knockout mutants of AUXIN BINDING PROTEIN 1 (ABP1), expecting them to fail to respond to auxin and have developmental defects, as previously seen in the abp1-1 knockdown mutant. Instead, these plants were indistinguishable from wild type plants, leading the authors to conclude that ABP1 is not required for auxin signaling or Arabidopsis development as previously believed.
Read the paper in PNAS: Auxin binding protein 1 (ABP1) is not required for either auxin signaling or Arabidopsis development.
A paper later in the year from the same authors found that the embryonic lethality of the abp1-1 mutant is actually caused by the off-target linked deletion of the adjacent BSM gene.
Read this paper in Nature Plants: Embryonic lethality of Arabidopsis abp1-1 is caused by deletion of the adjacent BSM gene.
The tale of ABP1 was examined in more detail on the GARNet blog, Weeding the Gems, which concluded: “In many ways this story is an excellent example of how science should work, where claims are independently tested to ensure that earlier experiments have been conducted or interpreted correctly.” Click here to read more.
A clever experiment from Germany led to a significant breakthrough in crop protection from insect pests.
When double-stranded RNA (dsRNA) is present within a eukaryotic cell, it is cleaved by the Dicer enzyme to form short interfering RNAs. These can bind to complementary RNA within a cell to target it for destruction, thus silencing the corresponding gene expression. This process is known as RNA interference (RNAi).
RNAi has previously been used to tackle insect herbivory by expressing insect-specific dsRNA in plants; however the protection has previously been incomplete. In this new study, published in Science, researchers produced dsRNA within chloroplasts, which do not have RNAi machinery. When dsRNA is expressed in the cytoplasm, the plant’s own Dicer enzyme breaks most of it down. When expressed in the chloroplasts, the dsRNA remained intact when eaten by insects, which proved much more effective at killing these pests.
Read the paper here: Full crop protection from an insect pest by expression of long double-stranded RNAs in plastids.
Another crop protection study followed in March, when researchers in China cloned the genetic locus in rice that confers broad-spectrum resistance to planthoppers – insect pests that cause the loss of billions of dollars of crops per year. Three lectin receptor kinase genes were found in rice cultivars from the Philippines, which enable plants to survive an infestation of insects. When cloned into a susceptible rice cultivar, these genes conferred resistance to two different planthopper species.
Understanding the genetic basis of resistance is very important as marker-assisted breeding and selection could be used to develop resistant rice varieties, and potentially utilized in other species of cereal.
Read the paper in Nature Biotechnology: A gene cluster encoding lectin receptor kinases confers broad-spectrum and durable insect resistance in rice
A European collaboration led to the development of 3DCellAtlas, a computational approach that semi-automatically identifies cell types in a developing 3D organ without the need for transgenic lineage markers. This program will enable the interpretation of dynamic organ growth and the spatial and temporal context of developmental cell divisions that produce the resultant plant. It could be integrated with growth in different conditions or with developmental mutants to examine exactly how these processes affect growth in 3D.
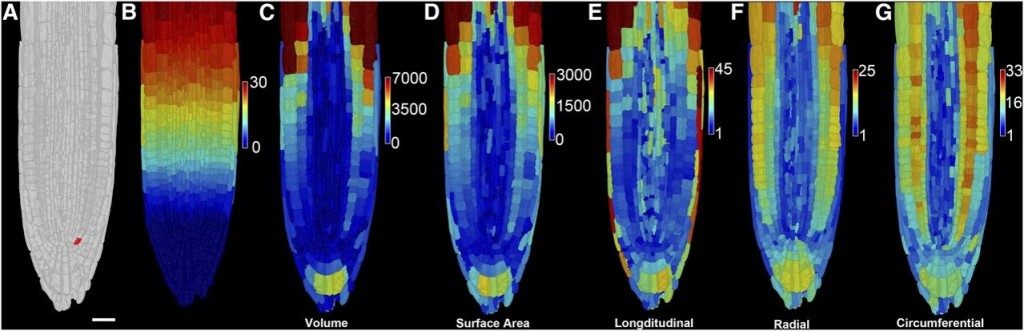
Image credit: Montenegro-Johnson et al., 2015. Digital Single-Cell Analysis of Plant Organ Development Using 3DCellAtlas. The Plant Cell, vol. 27 no. 4, 1018–1033.
A special issue of the Plant Biotechnology Journal was published in May, focusing on the amazing advances in molecular farming. While the entire issue is worth delving into, we were particularly intrigued by the review on moss-made pharmaceuticals, which outlines the rapid progress made in the field.
The model moss Physcomitrella patens has rapidly become one of the organisms of choice in biotechnology, with a fully sequenced genome and an outstanding toolbox for genome-engineering. The authors describe how moss-made pharmaceuticals can easily be produced while remaining remarkably more stable from batch to batch than cultured animal cells. The system is easily scalable, making their production highly cost effective, and safe. The first moss-made pharmaceuticals are currently in clinical trials, so keep an eye out for much more from this field over the next few years.
Read the review: Moss-made pharmaceuticals: from bench to bedside.
In June, US researchers discovered a new role for chloroplast stromules, protrusions that extend from the surface of all plastid types. The function of stromules has been difficult to determine, but this research, published in Developmental Cell, suggests that they may provide a mechanism by which plastid signals are conveyed to the nucleus. The paper shows that chloroplast stromules are induced by defense responses such as programmed cell death signaling, and that the stromules extend to form dynamic connections with the nucleus. The stromules may therefore aid in the amplification and/or transport of immune response signals into the nucleus.
Read the paper: Chloroplast Stromules Function during Innate Immunity.
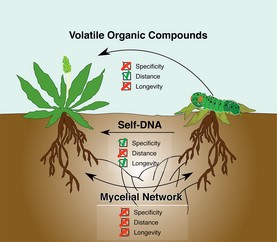
Image credit: Veresoglou et al., 2015. Self-DNA: a blessing in disguise? New Phytologist, vol. 207, no. 3, 488–490.
In late 2014 and early 2015, Italian researchers published a set of articles showing that extracellular self-DNA, DNA from conspecifics, could inhibit the growth of organisms from a wide range of taxa, including plants, bacteria, fungi and animals. Conversely, these organisms were not affected by extracellular DNA from other unrelated species.
In July, New Phytologist published a letter offering an interpretation of the data as it relates to plants. Plants could interpret extracellular self-DNA as an indicator of intraspecific competition (which seeds could use as a cue to remain dormant) or of a hostile environment that has already caused the death of conspecifics, signaling them to ramp up their pre-emptive immune response to increase survival after neighbors have been damaged or killed. There are still a lot of mechanisms and ecological effects to be investigated in this new field, but this letter suggests several interesting avenues to investigate.
Read the article: Self-DNA: a blessing in disguise?
Original research papers in New Phytologist:
Inhibitory effects of extracellular self-DNA: a general biological process?
A US study in August revealed a surprising degree of conservation in gene expression patterns across a wide range of plant taxa during root development. This was particularly interesting because the spikemoss Selaginella was shown to use many of the same genes as the evolutionarily distant angiosperms, despite the fossil record suggesting that roots evolved independently in these two lineages. Perhaps roots in these two groups evolved by independently recruiting the same developmental program, or perhaps by elaborating on a previously unknown proto-root that existed in the common ancestor of vascular plants.
Read the paper in The Plant Cell: Conserved Gene Expression Programs in Developing Roots from Diverse Plants.
Salt stress can significantly reduce the growth and yield of plants. Researchers in Germany identified two components of the cellulose synthase complex that directly interact with the microtubules and promote their dynamics, which interestingly were highly produced during salt stress conditions. During salt stress, cellulose microtubules depolymerize, however the newly discovered compounds, known as Companions of Cellulose Synthase, promote the reassembly of the microtubule to allow cellulose synthesis to continue.
Read the paper in Cell: A Mechanism for Sustained Cellulose Synthesis during Salt Stress
Throughout the year the GM debate in Europe reached several important milestones. In January the European Union (EU) changed its rules, giving individual countries more flexibility to decide for themselves whether or not to plant GM crops. In February, the UK Science and Technology Committee report stated that EU regulations preventing GM crops are not fit for purpose, and that they should be replaced with a trait-based system.
In October, EU member states revealed their stances on GM crops, with over half of Europe opting out of growing GM crops. Germany was the largest country to opt out of growing GM. The full list can be viewed here: Restrictions of geographical scope of GMO.
Read the news articles here:
EU changes rules on GM crop cultivation – January 2015
EU regulation on GM Organisms not ‘fit for purpose’ – February 2015
Half of Europe opts out of new GM crop scheme – October 2015

Image credit: University of Warwick, UK.
A collaboration between South African and UK scientists revealed how plants can use their circadian clock to pre-emptively boost their immune resistance at dawn, when fungal infection is most likely. Plants tend to decrease in susceptibility at dawn, but those with dysfunctional circadian clocks remained highly susceptible throughout the day. The research also showed that jasmonate signaling plays a crucial role in the circadian timing of resistance.
Read the article in The Plant Journal: Jasmonate signalling drives time-of-day differences in susceptibility of Arabidopsis to the fungal pathogen Botrytis cinerea.

Image credit: Guo & Liu., 2015. A single-nucleotide exon found in Arabidopsis. Scientific Reports, 5:18087.
Researchers in China published the surprising finding that a single-nucleotide exon exists in the APC11 gene in Arabidopsis. This is the smallest exon ever to be discovered before. The team used an elegant set of APC11-GFP constructs to show that intron splicing around the single-nucleotide exon is effective in both Arabidopsis and rice. This finding has implications for future genome annotations, which might reveal many more single-nucleotide exons.
Read the paper in Scientific Reports: A single-nucleotide exon found in Arabidopsis.
What a wonderful year of science! What new knowledge will 2016 bring?