The central biocatalyst in photosynthesis, Rubisco, is the most abundant enzyme on earth. By reconstructing billion-year-old enzymes, a team of Max Planck Researchers has deciphered one of the key adaptations of early photosynthesis. Their results not only provide insights into the evolution of modern photosynthesis but also offer new impulses for improving it.
Present day life fully depends on photosynthetic organisms like plants and algae that capture and convert CO2. At the heart of these processes lies an enzyme called Rubisco that captures more than 400 billion tons CO2 annually. Organisms alive today make staggering amounts of it: the mass of Rubisco on our planet outweighs that of all humans. In order to assume such a dominant role in the global carbon cycle, Rubisco had to adapt constantly to changing environmental conditions.
Using a combination of computational and synthetic approaches, a team from the Max-Planck-Institute for Terrestrial Microbiology in Marburg, Germany in collaboration with the University of Singapore has now successfully resurrected and studied billion-year-old enzymes in the lab. In this process, which they describe as “molecular paleontology”, the researchers found that instead of direct mutations in the active center, an entirely new component prepared photosynthesis to adapt to rising oxygen levels.
Rubisco’s early confusion
Rubisco is ancient: it emerged approximately four billion years ago in primordial metabolism prior to the presence of oxygen on earth. However, with the invention of oxygen-producing photosynthesis and rise of oxygen in the atmosphere, the enzyme started catalyzing an undesired reaction, in which it mistakes O2 for CO2 and produces metabolites that are toxic to the cell. This confused substrate scope still scars Rubiscos to date and limits photosynthetic efficiency. Even though Rubiscos that evolved in oxygen-containing environments became more specific for CO2 over time, none of them could get completely rid of the oxygen capturing reaction.
The molecular determinants of increased CO2 specificity in Rubisco remain largely unknown. However, they are of great interest to researchers aiming to improve photosynthesis. Interestingly, those Rubiscos that show increased CO2specificity recruited a novel protein component of unknown function. This component was suspected to be involved in increasing CO2 specificity, however, the true reason for its emergence remained difficult to determine because it already evolved billions of years ago.
Studying evolution by resurrecting ancient proteins in the lab
To understand this key event in the evolution of more specific Rubiscos, collaborators at the Max Planck Institute for Terrestrial Microbiology in Marburg and Nanyang Technological University in Singapore used a statistical algorithm to recreate forms of Rubiscos that existed billions of years ago, before oxygen levels began to rise. The team led by Max Planck researchers Tobias Erb and Georg Hochberg resurrected these ancient proteins in the lab to study their properties. In particular, the scientists wondered whether Rubisco’s new component had anything to do with the evolution of higher specificity.
The answer was surprising, as doctoral researcher Luca Schulz explains: “We expected the new component to somehow directly exclude oxygen from Rubisco catalytic center. That is not what happened. Instead, this new subunit seems to act as a modulator for evolution: recruitment of the subunit changed the effect that subsequent mutations had on Rubisco’s catalytic subunit. Previously inconsequential mutations suddenly had a huge effect on specificity when this new component was present. It seems that having this new subunit completely changed Rubisco’s evolutionary potential.”
An enzyme’s addiction to its new subunit
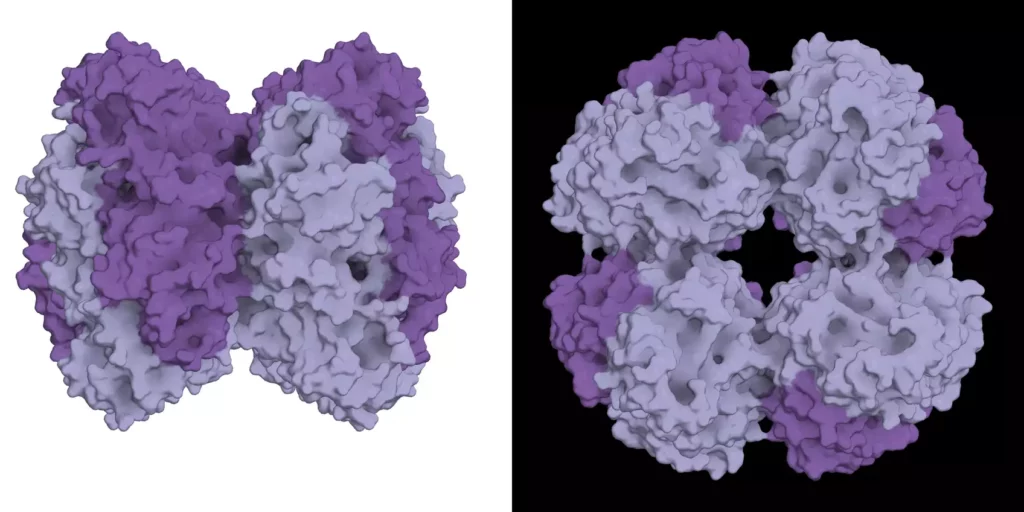
This function as an “evolutionary modulator” also explains another mysterious aspect of the new protein component: Rubiscos that incorporated it are completely dependent on it, even though other forms of Rubisco can function perfectly well without. The same modulating effect explains why: When bound to this small protein component, Rubisco become tolerant to mutations that would otherwise be catastrophically detrimental. With the accumulation of such mutations, Rubisco effectively became addicted to its new subunit.
Altogether the findings finally explain the reason why Rubisco kept this new protein component around ever since it encountered it. Max Planck Research Group Leader Georg Hochberg explains: “The fact that this connection was not understood until now highlights the importance of evolutionary analysis for understanding the biochemistry that drives life around us. The history of biomolecules like Rubisco can teach us so much about why they are the way they are today. And there are still so many biochemical phenomena whose evolutionary history we really have no idea about. So it’s a very exciting time to be an evolutionary biochemist: almost the entire molecular history of the cell is still waiting to be discovered.”
Scientific journeys back in time can provide invaluable insights for the future
The study also has important implications for how photosynthesis might be improved, says Max Planck Director Tobias Erb: “Our research taught us that traditional attempts to improve Rubisco might have been looking in the wrong place: for years, research focused solely on changing amino acids in Rubisco itself to improve it. Our work now suggests that adding entirely new protein components to the enzyme could be more productive and may open otherwise impossible evolutionary paths. This is uncharted land for enzyme engineering.”
Read the paper: Science
Article source: Max Planck Society
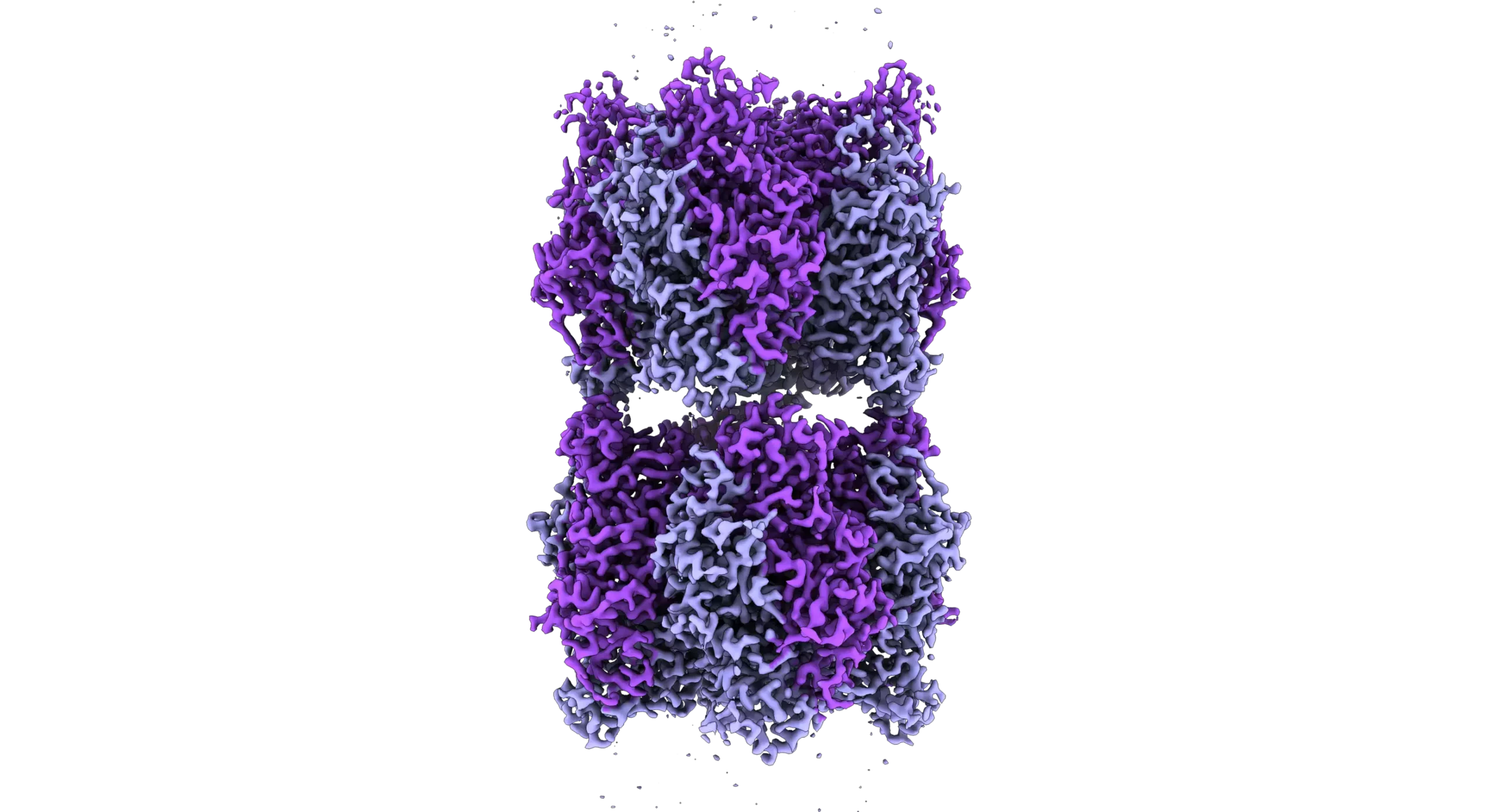
Image: Cryo-electron microscope image of two Rubisco complexes interacting with each other. If a subunit essential for solubility is missing, individual enzyme complexes can interact with each other in this way and form thread-like structures, so-called fibrils. Under normal conditions, however, Rubisco does not form such fibrils. Credit: MPI f. Terrestrial Microbiology/ L. Schulz